Dynamic genetic analysis for body weight and main length ratio in turbot Scophthalmus maximus
-
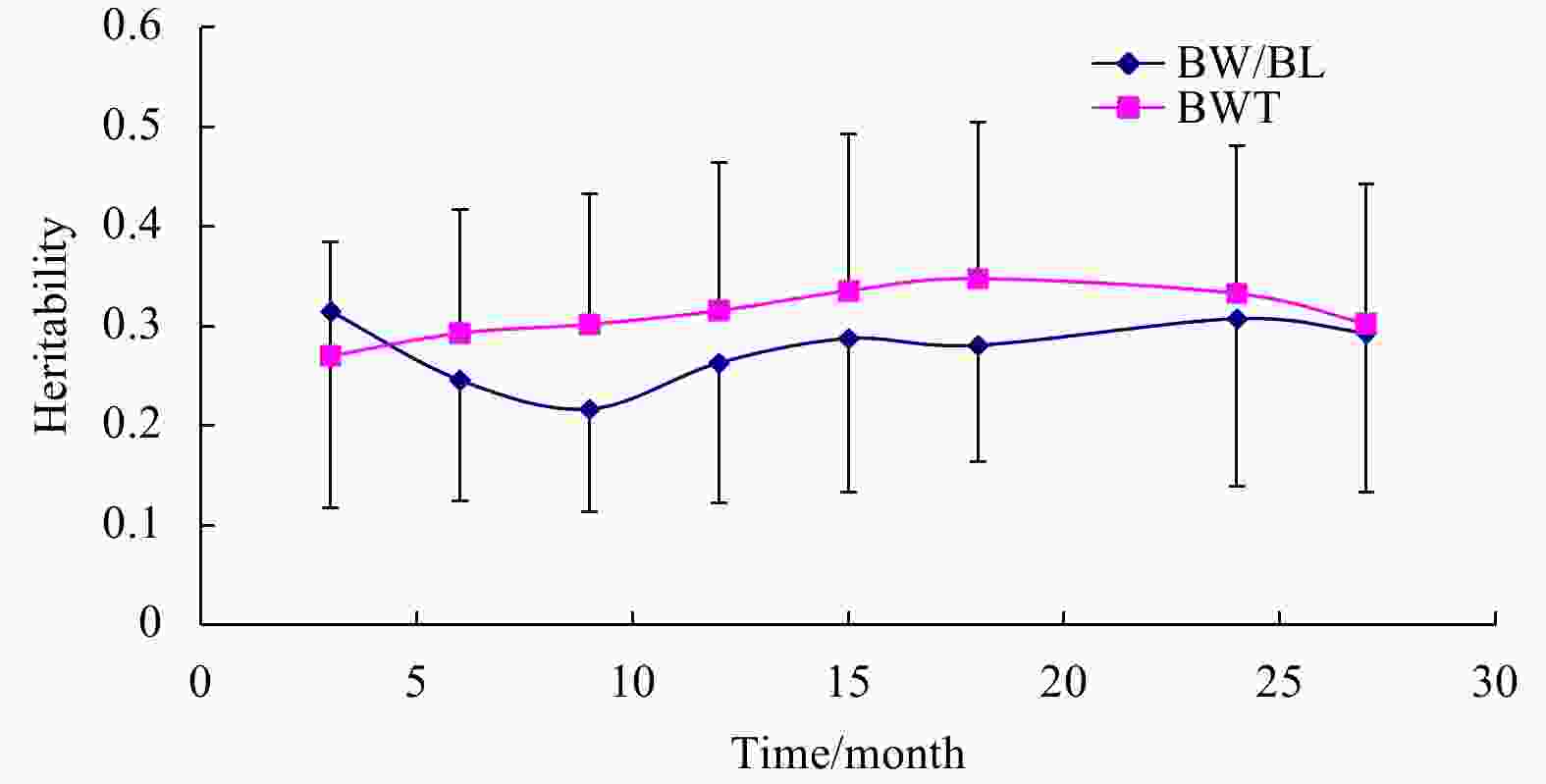
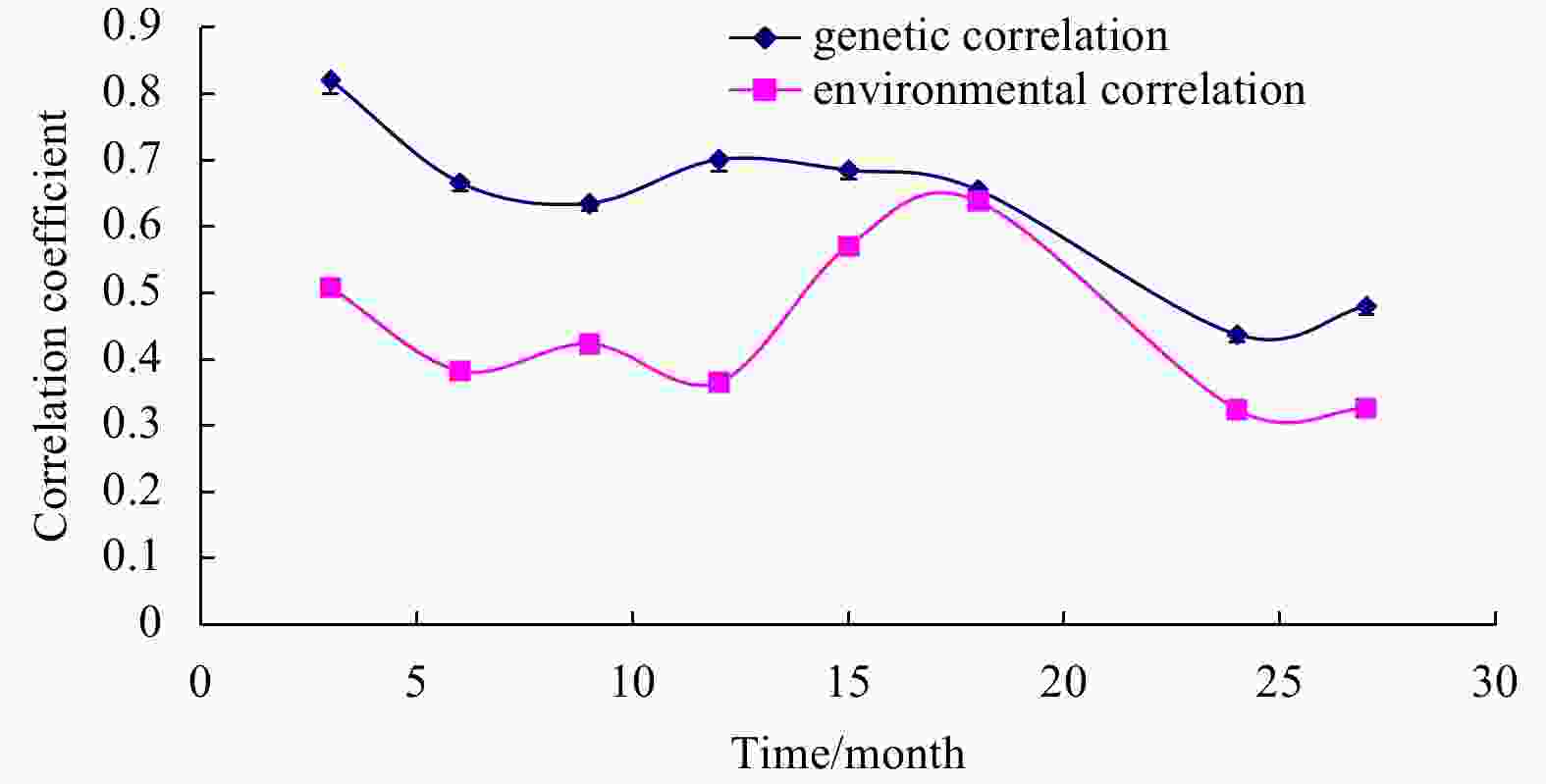
Abstract: The objective of this study was to estimate genetic parameters of body width (BW) to body length (BL) ratio (BW/BL) and of body weight traits (BWT) in turbot, and to elucidate the genetic mechanism of the two traits during ontogeny by dynamic genetic analysis. From 3 to 27 months, BW, BL and BWT of each communally stocked fish were measured every 3 months. The BW/BL ratio was measured at different sampling ages. A two-trait animal model was used for genetic evaluation of traits. The results showed that the heritability values of BW/BL ratio ranged from 0.216 8 to 0.314 8, corresponding to moderate heritability. The BWT heritability values ranged from 0.270 2 to 0.347 9 corresponding to moderate heritability. The heritability of BW/BL ratio was lower than that of BWT, except at 3 months of age. Genetic correlation between BW/BL ratio and BWT decreased throughout the measurement period. Genetic correlations were higher than the phenotypic correlations. The current results for estimating genetic parameters demonstrate that the BW/BL ratio could be used as a phenotypic marker of fast-growing turbot, and the BW/BL ratio and BWT could be improved simultaneously through selective breeding.
-
Key words:
- turbot /
- dynamic genetic analysis /
- body weight /
- main length ratio /
- heritability /
- genetic correlation
-
Table 1. Ratio of body width/body length (BW/BL ratio), and mean body weight (BWT) for each family of turbot at different sampling ages (mean±standard deviation)
Months of age BW/BL ratio BWT/g 3 0.605 6±0.022 9 2.979 1±0.916 2 6 0.784 4±0.025 0 31.125 0±9.181 8 9 0.786 1±0.024 2 164.718 8±21.533 3 12 0.813 0±0.034 5 376.412 0±47.118 2 15 0.795 4±0.0314 593.113 1±76.325 6 18 0.827 9±0.0572 2 996.683 4±107.853 4 24 0.819 3±0.048 9 1 776.437 8±183.442 1 27 0.856 7±0.332 1 2 031.383 2±279.890 1 Table 2. Variance components and heritability (h2) with standard errors (mean±SE) of BW/BL ratio of turbot at different sampling ages
Months of age $\sigma _a^2$ $\sigma _f^2$ $\sigma _e^2$ ${h^2}$ 3 0.150 0±0.003 8 0.112 8±0.000 1 0.213 7±0.002 7 0.314 8±0.197 1 6 0.122 9±0.012 6 0.083 4±0.000 3 0.293 6±0.013 5 0.245 8±0.121 5 9 0.256 3±0.017 1 0.100 0±0.002 4 0.825 9±0.057 8 0.216 8±0.103 2 12 1.265 2±0.135 9 0.130 0±0.018 4 3.413 4±0.179 3 0.263 1±0.140 6 15 5.580 3±1.274 9 0.089 0±0.001 3 13.729 5±2.458 1 0.287 6±0.154 3 18 18.720 2±3.946 6 0.100 0±0.003 5 47.807 3 ±7.073 4 0.280 9±0.117 2 24 27.350 8±5.001 2 0.095 1±0.001 4 61.501 1±10.713 0 0.307 5±0.168 3 27 33.331 1±8.701 1 0.016 0±0.001 1 80.312 1±12.673 2 0.293 3±0.159 8 Note: $\sigma _a^2$ represents additive genetic variance, $\sigma _f^2$ full-sib variance, $\sigma _e^2$ residual variance, h2 heritability, BW body width, and BL body length. Table 3. Variance components and heritability (h2) with standard errors (mean±SE) of BWT of turbot at different sampling ages
Months of age $\sigma _a^2$ $\sigma _f^2$ $\sigma _e^2$ ${h^2}$ 3 0.101 5±0.019 3 0.223 4±0.001 1 0.050 7±0.040 2 0.270 2±0.114 3 6 12.365 1±6.124 1 0.194 4±0.001 2 29.685 3±11.365 1 0.293 1±0.123 6 9 276.341 6±66.606 0 0.210 6±0.001 4 638.884 3±105.131 6 0.301 9±0.130 8 12 700.260 0±101.098 6 0.240 6±0.001 7 2 218.330 0±283.817 3 0.315 6±0.149 0 15 1 950.150 0±196.814 1 0.199 6±0.001 1 3 858.514 0±415.443 1 0.335 6±0.157 7 18 4 047.030 0±527.378 4 0.210 6±0.001 4 7 584.000 1±994.087 2 0.347 9±0.156 5 24 29 679.150 0±3 043.887 7 0.205 7±0.001 5 59 496.430 0±6 211.432 1 0.332 816±0.148 8 27 49 368.553 2±6 003.087 0 0.126 6±0.001 2 113 783.300 1±2 0583.076 3 0.302 5±0.140 1 Note: $\sigma _a^2$ represents additive genetic variance, $\sigma _f^2$full-sib variance, $\sigma _e^2$residual variance, h2 heritability, and BWT body weight. Table 4. Genetic and phenotypic correlations between BW/BL ratio and BWT
Months of age Genetic correlation (${r_{{A_1}{A_2}}}$) Phenotypic correlation (${r_{{P_1}{P_2}}}$) 3 0.821 3±0.021 6** 0.534 0±0.000 3** 6 0.666 7±0.013 7** 0.402 7±0.000 1** 9 0.635 5±0.012 1** 0.435 9±0.000 2** 12 0.701 4±0.017 8** 0.392 9±0.000 0** 15 0.685 4±0.014 2** 0.581 2±0.000 0** 18 0.655 5±0.011 9** 0.638 3±0.000 1** 24 0.437 8±0.012 2** 0.336 1±0.000 0** 27 0.480 4±0.012 9** 0.340 1±0.000 0** Note: * A significant correlation (P<0.05); ** a highly significant correlation (P<0.01). BW represents body width, BL body length, and BWT body weight. -
[1] Ankorion Y, Moav R, Wohlfarth G W. 1992. Bidirectional mass selection for body shape in common carp. Genetics Selection Evolution, 24: 43. doi: 10.1186/1297-9686-24-1-43 [2] Awan Z K, Masood S A, Naseem Z, et al. 2015. Genetic variability and trait association for relative growth rate and green fodder yield in Sorghum bicolorl. International Journal of Biology Pharmacy and Allied Sciences, 4(8): 5566–5576 [3] Battaglene S C. 1996. Hormone-induced ovulation of sand whiting (Sillago ciliate). Asian Fisheries Science, 9: 169–176 [4] Blonk R J W, Komen J, Tenghe A, et al. 2010. Heritability of shape in common sole, Solea solea, estimated from image analysis data. Aquaculture, 307(1–2): 6–11. doi: 10.1016/j.aquaculture.2010.06.025 [5] Cardellino R, Rovira J. 1987. Mejoramiento Genético Animal (in Spanish). Buenos Aires: Hemisferio Sur, 253 [6] De Oliveira C A L, Ribeiro R P, Yoshida G M, et al. 2016. Correlated changes in body shape after five generations of selection to improve growth rate in a breeding program for Nile tilapia Oreochromis niloticus in Brazil. Journal of Applied Genetics, 57(4): 487–493. doi: 10.1007/s13353-016-0338-5 [7] Gilmour A R, Gogel B J, Cullis B R, et al. 2009. ASReml User Guide Release 3.0. Hemel Hempstead, UK: VSN International Ltd [8] Gjerde B, Schaeffer L R. 1989. Body traits in rainbow trout: II. Estimates of heritabilities and of phenotypic and genetic correlations. Aquaculture, 80(1–2): 25–44. doi: 10.1016/0044-8486(89)90271-8 [9] Guan C T. 2016. Annual Report 2015 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 179–242 [10] Islam M S, Khan S, Khanam D, et al. 1993. Genetic variability and path analysis in cucumber (Cucumis sativus L.). Bangladesh Journal of Plant Breeding and Genetics, 6: 45–51 [11] Kause A, Ritola O, Paananen T, et al. 2003. Big and beautiful? Quantitative genetic parameters for appearance of large rainbow trout. Journal of Fish Biology, 62(3): 610–622. doi: 10.1046/j.1095-8649.2003.00051.x [12] Kirkpatrick M, Heckman N. 1989. A quantitative genetic model for growth, shape, reaction norms, and other infinite-dimensional characters. Journal of Mathematical Biology, 27(4): 429–450. doi: 10.1007/BF00290638 [13] Kirkpatrick M, Lofsvold D, Bulmer M. 1990. Analysis of the inheritance, selection and evolution of growth trajectories. Genetics, 124(4): 979–993 [14] Kumar V, Singh D K, Panchbhaiya A, et al. 2017. Correlation and path coefficient analysis studies in midseason cauliflower (Brassica oleracea var. botrytis L.). Journal of Pharmacognosy and Phytochemistry, 6(4): 1130–1137 [15] Lei Jilin. 2010. Annual Report 2009 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 75–120 [16] Lei Jilin. 2012. Annual Report 2011 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 117–176 [17] Lei Jilin. 2013. AnnuaL Report 2012 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 183–242 [18] Lei Jilin. 2014. Annual Report 2013 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 187–246 [19] Lei Jilin. 2015. AnnuAl Report 2014 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 199–266 [20] Liu Yongxin, Jiang Li, Liu Haijin, et al. 2014. Phenotypic and genetic parameter estimation of morphological traits related to axial body growth in Japanese flounder. Fisheries Science, 80(2): 317–321. doi: 10.1007/s12562-014-0704-3 [21] Liu Yongxin, Sun Zhaohui, Wang Yufen, et al. 2015. Genetic analysis for main length ratio associated with morphological traits in Japanese flounder Paralichthys olivaceus. Journal of Fish Biology, 86(3): 1129–1138. doi: 10.1111/jfb.12635 [22] Ma Aijun, Chen Chao, Lei Jilin, et al. 2006. Turbot Scophthalmus maximus: stocking density on growth, pigmentation and feed conversion. Chinese Journal of Oceanology and Limnology, 24(3): 307–312. doi: 10.1007/BF02842633 [23] National Technology Research and Development Center for Flounder Industry. 2011. AnnuaL Report 2010 of National Technology System for Flatfish Culture Industry (in Chinese). Qingdao: China Ocean University Press, 129–174 [24] Rana N P, Pandit M K. 2011. Studies on the genetic variability, character association and path analysis in snake gourd (Trichosanthes anguina L.) genotypes. Journal of Crop and Weed, 7(2): 91–96 [25] Ruan X H, Wang W J, Kong J, et al. 2011. Isolation and analysis of microsatellites in the genome of turbot (Scophthalmus maximus L.). African Journal of Biotechnology, 10(4): 507–518 [26] Saatci M, Dewi I A, Aksoy A R. 2003. Application of REML procedure to estimate the genetic parameters of weekly liveweights in one-to-one sire and dam pedigree recorded Japanese quail. Journal of Animal Breeding and Genetics, 120(1): 23–28. doi: 10.1046/j.1439-0388.2003.00370.x [27] Said E M, Fatiha H. 2015. Genotypic variation in fruit characters in some genotypes of watermelon cultivated in Morocco. International Journal of Agronomy and Agricultural Research, 6(4): 130–137 [28] Schaeffer L R. 2004. Application of random regression models in animal breeding. Livestock Production Science, 86(1–3): 35–45. doi: 10.1016/S0301-6226(03)00151-9 [29] Shikano T. 2007. Quantitative genetic parameters for growth-related and morphometric traits of hatchery-produced Japanese flounder Paralichthys olivaceus in the wild. Aquaculture Research, 38(12): 1248–1253. doi: 10.1111/j.1365-2109.2007.01749.x [30] Shimada Y, Shikano T, Murakami N, et al. 2007. Maternal and genetic effects on individual variation during early development in Japanese flounder Paralichthys olivaceus. Fisheries Science, 73(2): 244–249. doi: 10.1111/j.1444-2906.2007.01330.x [31] Wang Xin’an, Ma Aijun. 2015. Comparison of the morphometric dynamics of fast-growing and slow-growing strains of turbot Scophthalmus maximus. Chinese Journal of Oceanology and Limnology, 33(4): 890–894. doi: 10.1007/s00343-015-4195-6 [32] Wang Xin’an, Ma Aijun. 2016. Comparison of four nonlinear growth models for effective exploration of growth characteristics of turbot Scophthalmus maximus fish strain. African Journal of Biotechnology, 15(40): 2251–2258. doi: 10.5897/AJB2016.15490 [33] Wang Xin’an, Ma Aijun, Huang Zhihui, et al. 2010. Heritability and genetic correlation of survival in turbot (Scophthalmus maximus). Chinese Journal of Oceanology and Limnology, 28(6): 1200–1205. doi: 10.1007/s00343-010-9014-5 [34] Wang Xin’an, Ma Aijun, Ma Deyou. 2015. Developmental quantitative genetic analysis of body weights and morphological traits in the turbot, Scophthalmus maximus. Acta Oceanologica Sinica, 34(2): 55–62. doi: 10.1007/s13131-015-0618-7 [35] Xu Liyong, Wang Weiji, Kong Jie, et al. 2015. Estimates of heritability and correlation for growth traits of Turbot (Scophthalmus maximus L.) under low temperature conditions. Acta Oceanologica Sinica, 34(2): 63–67. doi: 10.1007/s13131-015-0616-9 [36] Zhang Tianshi, Kong Jie, Liu Baosuo, et al. 2014. Genetic parameter estimation for juvenile growth and upper thermal tolerance in turbot (Scophthalmus maximus Linnaeus). Acta Oceanologica Sinica, 33(8): 106–110. doi: 10.1007/s13131-014-0460-3 -




 下载:
下载: