A new eyeless species of Nicon (Annelida: Nereididae) from the deep Northwest Pacific Ocean
-
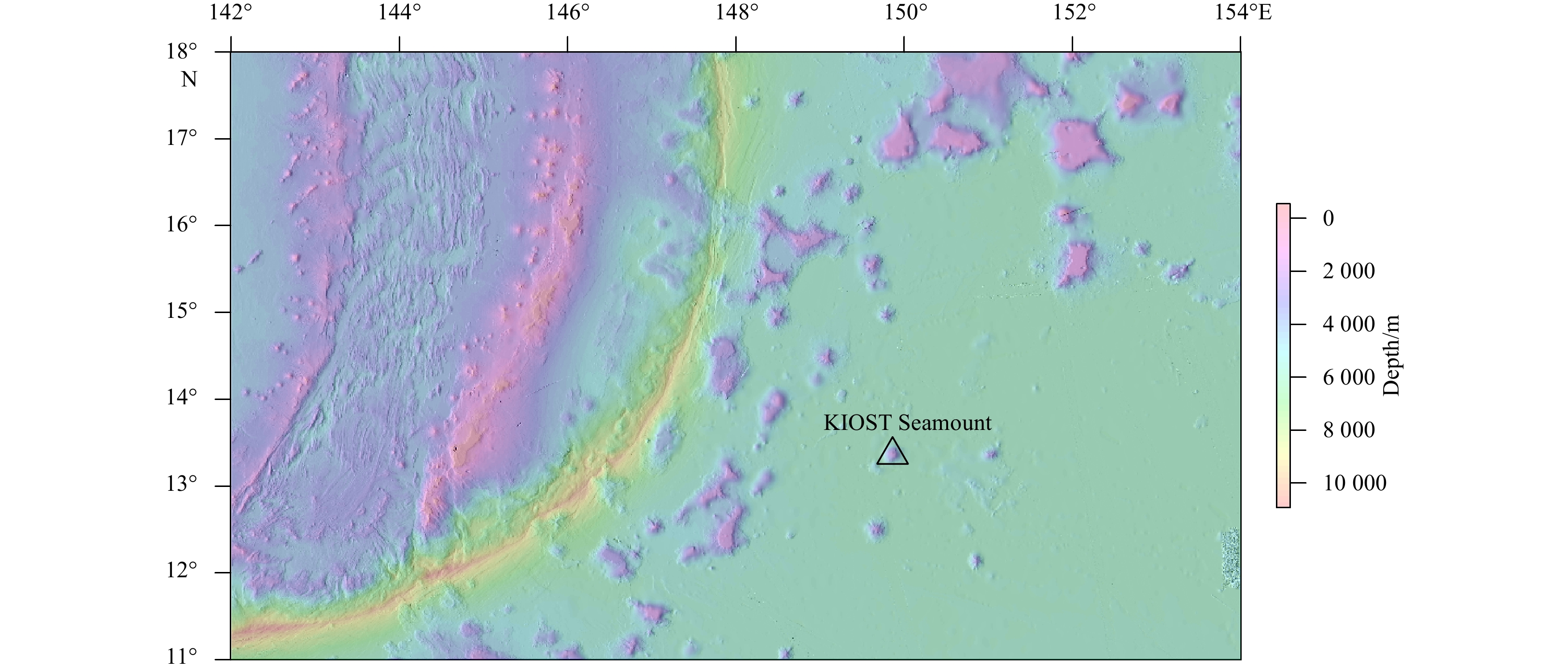
Abstract: A new species of the nereidid annelid, genus Nicon Kinberg, 1866, from KIOST Seamount, Northwest Pacific deep water is described. Nicon is a genus characterized by lacking paragnaths or papillae on the pharynx and composed of nine species worldwide, distributed from shallow water to deep sea. Nicon ablepsia sp. nov. here described is characterized by the lack of eyes on the prostomium, prolonged tentacular cirri reaching to chaetiger 6, notochaetae homogomph spinigers, neurochaetae homogomph spinigers and heterogomph falcigers. Phylogenetic relationships of Nicon remain undetermined based on molecular data. In this study, we constructed molecular Maximum-Likelihood phylogenetic tree from 29 nereidid species based on four marker genes: mitochondrial 16S rRNA gene and cytochrome c oxidase subunit I (COI) gene; nuclear 18S rRNA gene and 28S rRNA gene. Our analysis suggest the Nicon is clustered within Nereidinae, and nereidinae is not recovered as monophyletic. A key to species of Nicon is provided.
-
Key words:
- deep sea /
- Nicon /
- new species /
- Gymnonereidinae /
- systematics
-
Figure 2. Nicon ablepsia sp. nov.: a. dorsal view of the holotype; b. lateral view of the holotype, white triangles mark tentacular cirri; c. dorsal view of head; d. ventral view of head; e. jaws; f. chaetigers 8–17, dorsal view; g. chaetiger 7, end view; and h. chaetiger 7, frontal view. Scale bars: 2.0 mm (a), 0.5 mm (e), 0.05 mm (b, d, f, h).
Figure 3. Parapodia and chaetae (a−d, SEM; e−g, optical picture). a. Parapodium; b. notopodial lobe, arrow shows curved tip of notoacicula; c. neurochaetae, homogomph spinigers and heterogomph falcigers; d. notochaetae, homogomph spinigers; e. neuropodium with neurochaetae; f. heterogomph falcigers; and g. notochaetae, homogomph spinigers. Scale bars: 0.5 mm (a), 0.1 mm (b−d).
Figure 4. The Maximum-Likelihood phylogenetic tree of 29 species based on the concatenated dataset of 16S rDNA, COI gene, 18S rDNA and 28S rDNA sequences with 1 000 bootstrap replicates. Bootstrap support values (calculated by RAxML software) are indicated adjacent to each node. The red solid dot shows the inconsistent topology cluster between the tree constructed by RAxML and IQ-TREE. All the other nodes were identical.
A1. The Maximum-Likelihood tree constructed by IQ-TREE. Numbers near the branch nodes refer to SH-aLRT/ UFBoot support values (all based on 1 000 replicates). The red solid dot showed the inconsistent topology cluster between the tree constructed by RAxML and IQ-TREE. The partitioned scheme and best model information was auto selected by IQ-TREE as follows: Part 1 (1–1 158 nt, 16S rDNA), GTR+F+I+G4 model; Part 2 (1 159–2 697 nt, COI gene), TIM2+F+R4 model; Part 3 (2 698–4 600 nt, 18S rDNA), TIM2e+FQ+I+G4 model; and Part 4 (4 601–5 559 nt, 28S rDNA), TN+F model.
Table 1. Information and GenBank accession No. of marker genes used in the phylogenetic analysis
Subfamily/family Taxa 16S rDNA, length/bp COI gene, length/bp 18S rDNA, length/bp 28S rDNA, length/bp Nereidinae Alitta succinea MN823959, 352 MN823952, 1 515 AY210447, 1 891 AY210464, 3 504 Ceratonereis longiceratophora − AY583701, 400 AB106251, 1 642 AF185189, 533 Hediste japonica LC323064, 416 LC323029, 570 − LC380658, 804 Hediste diadroma KX499500, 1 196 KX499500, 1 535 LC323646, 1 698 LC380656, 804 Nectoneanthes oxypoda − MN256616, 658 KX290701, 1 793 LC168841, 527 Nereis heterocirrata KC833492, 462 MN256591, 658 KC840697, 790 − Nereis pelagica AY340470, 465 HQ024126, 660 AY340438, 1 795 − Nereis vexillosa GU362677, 442 MF121661, 658 DQ790083, 1 829 DQ790043, 3 087 Nereis sp. MF960765, 1 182 MF960765, 1 534 − − Paraleonnates uschakovi KX462988, 1 191 KX462988, 1 533 − − Perinereis wilsoni LC482173, 422 MN256542, 658 KC840691, 779 − Perinereis aibuhitensis KF611806, 1 197 KF611806, 1 534 − − Perinereis cultrifera MN812983, 1 016 MN812983, 1 534 − − Perinereis nuntia JX644015, 1 199 JX644015, 1 537 − − Perinereis sp. MN823971, 1 138 MN823962, 1 536 − − Platynereis cf. australis MN830369, 1 196 MN830369, 1 534 − − Platynereis bicanaliculata MN812984, 1170 MN812984, 1 534 − − Platynereis dumerilii AF178678, 1172 AF178678, 1 534 AY894303, 1 805 − Pseudonereis variegata MN855213, 613 MN855134, 1308 KC840693, 779 − Gymnonereidinae Ceratocephale abyssorum GQ426618, 493 GQ426683, 421 GQ426585, 1 709 − Gymnonereis sp. KY704332, 472 KY805814, 633 − − Laeonereis culveri KU992689, 1 007 KU992689, 1 536 − − Tylorrhynchus heterochaetus KM111507, 1 236 KM111507, 1 534 − − Nicon ablepsia sp. nov. MW525220, 513 MW644966, 1 534 MW525221, 1 820 MW525222, 1 059 Namanereidinae Namalycastis abiuma KU351089, 1 019 KU351089, 1 534 − − Namalycastis hawaiiensis LC213728, 521 MN125542, 658 LC213729, 1 781 LC213727, 816 Namalycastis indica MF959005, 482 MF958995, 660 − MF959019, 766 Namalycastis jaya JX483870, 462 JN790067, 699 JX483866, 1 744 − Hesionidae Vrijenhoekia balaenophila JN571884, 512 JN571831, 658 JN571895, 1 777 JN571904, 781 Note: − means the sequence was not reported. The text in bold indicates the measurement results in this paper. -
[1] Alves P R, Halanych K M, Santos C S G. 2020. The phylogeny of Nereididae (Annelida) based on mitochondrial genomes. Zoologica Scripta, 49(3): 366–378. doi: 10.1111/zsc.12413 [2] Bakken T, Wilson R S. 2005. Phylogeny of nereidids (Polychaeta, Nereididae) with paragnaths. Zoologica Scripta, 34(5): 507–547. doi: 10.1111/j.1463-6409.2005.00200.x [3] Bankevich A, Nurk S, Antipov D, et al. 2012. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol, 19(5): 455–477. doi: 10.1089/cmb.2012.0021 [4] Bonyadi-Naeini A, Rastegar-Pouyani N, Rastegar-Pouyani E, et al. 2017. Nereididae (Annelida: Phyllodocida) of the Persian Gulf and Gulf of Oman, including description of two new species and 11 new records. Zootaxa, 4244(1): 91–117. doi: 10.11646/zootaxa.4244.1.5 [5] Chernomor O, von Haeseler A, Minh B Q. 2016. Terrace aware data structure for phylogenomic inference from supermatrices. Systematic Biology, 65(6): 997–1008. doi: 10.1093/sysbio/syw037 [6] Conde-Vela V M, Wu Xuwen. 2019. Revision of Sinonereis Wu & Sun, 1979 (Annelida: Nereididae). Zoosystema, 41(1): 153–161. doi: 10.5252/zoosystema2019v4a9 [7] Conde-Vela V M, Wu Xuwen, Salazar-Vallejo S I. 2018. Reevaluation and new species of Kainonereis Chamberlin, 1919 (Annelida: Polychaeta: Nereididae). Zoological Studies, 57: e6 [8] de León-González J A, Trovant B. 2013. A new species of Nicon Kinberg, 1866 (Polychaeta, Nereididae) from Ecuador, Eastern Pacific, with a key to all known species of the genus. ZooKeys, 269: 67–76. doi: 10.3897/zookeys.269.4003 [9] Fauchald K. 1972. Benthic polychaetous annelids from deep water off western Mexico and adjacent areas in the eastern Pacific Ocean. Allan Hancock Monographs in Marine Biology, 7: 1–575 [10] Fitzhugh K. 1987. Phylogenetic relationships within the Nereididae (Polychaeta): implications at the subfamily level. Bulletin of the Biological Society of Washington, 7: 174–183 [11] Gonzalez B C, Martínez A, Borda E, et al. 2018. Phylogeny and systematics of Aphroditiformia. Cladistics, 34(3): 225–259. doi: 10.1111/cla.12202 [12] Hartman O. 1960. Systematic account of some marine invertebrate animals from the deep basins off southern California. Allan Hancock Pacific Expeditions, 22: 69–216 [13] Hartman O. 1967. Polychaetous annelids collected by the USNS Eltanin and Staten Island cruises, chiefly from Antarctic seas. Allan Hancock Monographs in Marine Biology, 2: 1–387 [14] Hartman O, Fauchald K. 1971. Deep-water benthic polychaetous annelids off New England to Bermuda and other north Atlantic areas Part II. Allan Hancock Monographs in Marine Biology, 6: 1–327 [15] Kirkegaard J B. 1995. Bathyal and abyssal polychaetes (errant species). Galathea Report, 17: 7–56 [16] Lanave C, Preparata G, Saccone C, et al. 1984. A new method for calculating evolutionary substitution rates. Journal of Molecular Evolution, 20(1): 86–93. doi: 10.1007/BF02101990 [17] Monro C C A. 1930. Polychaete worms. Discovery Reports, 2: 1–222 [18] Nguyen L T, Schmidt H A, von Haeseler A, et al. 2015. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution, 32(1): 268–274. doi: 10.1093/molbev/msu300 [19] Norlinder E, Nygren A, Wiklund H, et al. 2012. Phylogeny of scale-worms (Aphroditiformia, Annelida), assessed from 18SrRNA, 28SrRNA, 16SrRNA, mitochondrial cytochrome c oxidase subunit I (COI), and morphology. Molecular Phylogenetics and Evolution, 65(2): 490–500. doi: 10.1016/j.ympev.2012.07.002 [20] Paterson G L J, Glover A G, Barrio Froján C R S, et al. 2009. A census of abyssal polychaetes. Deep Sea Research Part II: Topical Studies in Oceanography, 56(19−20): 1739–1746. doi: 10.1016/j.dsr2.2009.05.018 [21] Santos C S G, Pleijel F, Lana P, et al. 2005. Phylogenetic relationships within Nereididae (Annelida: Phyllodocida). Invertebrate Systematics, 19(6): 557–576. doi: 10.1071/IS05001 [22] Shimabukuro M, Santos C S G, Alfaro-Lucas J M, et al. 2017. A new eyeless species of Neanthes (Annelida: Nereididae) associated with a whale-fall community from the deep Southwest Atlantic Ocean. Deep Sea Research Part II: Topical Studies in Oceanography, 146: 27–34. doi: 10.1016/j.dsr2.2017.10.013 [23] Silvestro D, Michalak I. 2012. raxmlGUI: a graphical front-end for RAxML. Organisms Diversity & Evolution, 12(4): 335–337 [24] Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30(9): 1312–1313. doi: 10.1093/bioinformatics/btu033 [25] Yang Ziheng. 1994. Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. Journal of Molecular Evolution, 39(3): 306–314. doi: 10.1007/BF00160154 [26] Zhang Yanjie, Sun Jin, Chen Chong, et al. 2017. Adaptation and evolution of deep-sea scale worms (Annelida: Polynoidae): insights from transcriptome comparison with a shallow-water species. Scientific Reports, 7(1): 46205. doi: 10.1038/srep46205 -




 下载:
下载: