Molecular quantification of copepod Acartia erythraea feeding on different algae preys
-
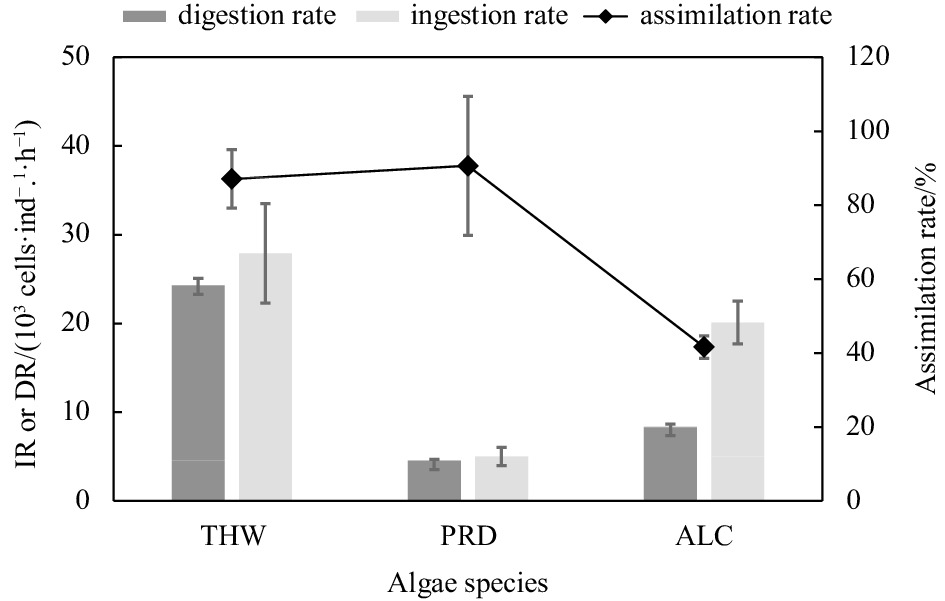
Abstract: Quantitative evaluation of the copepod feeding process is critical for understanding the functioning of marine food webs, as this provides a major link between primary producers and higher trophic levels. In this study, a molecular protocol based on quantitative polymerase chain reaction (qPCR) targeting 18S rDNA was developed and used to investigate the feeding and digestion rates of the copepod Acartia erythraea in a laboratory experiment using microalgae Thalassiosira weissflogii, Prorocentrum shikokuense, and Alexandrium catenella as prey. Although offered an equal encounter rate based on biovolume, prey uptake varied substantially among the three algal species, with the ingestion rate (IR) and digestion rate (DR) of A. erythraea differing significantly (P< 0.001) based on both cell counting and qPCR detection.Acartia erythraea showed the highest IR (2.79 × 104 cells/(ind.·h)) and DR (2.43 × 104 cells/(ind.·h)) on T. weissflogii, and the lowest amounts of ingested P. shikokuense were detected. The highest assimilation rate (~90.64%, IR/DR) was observed in copepods fed with P. shikokuense. The qPCR method used here can help determine the digestion rate and assimilation rate of copepods by detecting cells remaining in the gut hence providing the possibility to examine trophic links involving key species in the marine ecosystem. Our results indicate that A. erythraea has diet-specific feeding performance in different processes, and a quantitative assessment of copepod feeding is needed to accurately determine its functional role in the energy and matter uptake from marine food webs.
-
Key words:
- copepod /
- ingestion rate /
- digestion rate /
- 18S rDNA /
- real-time PCR
-
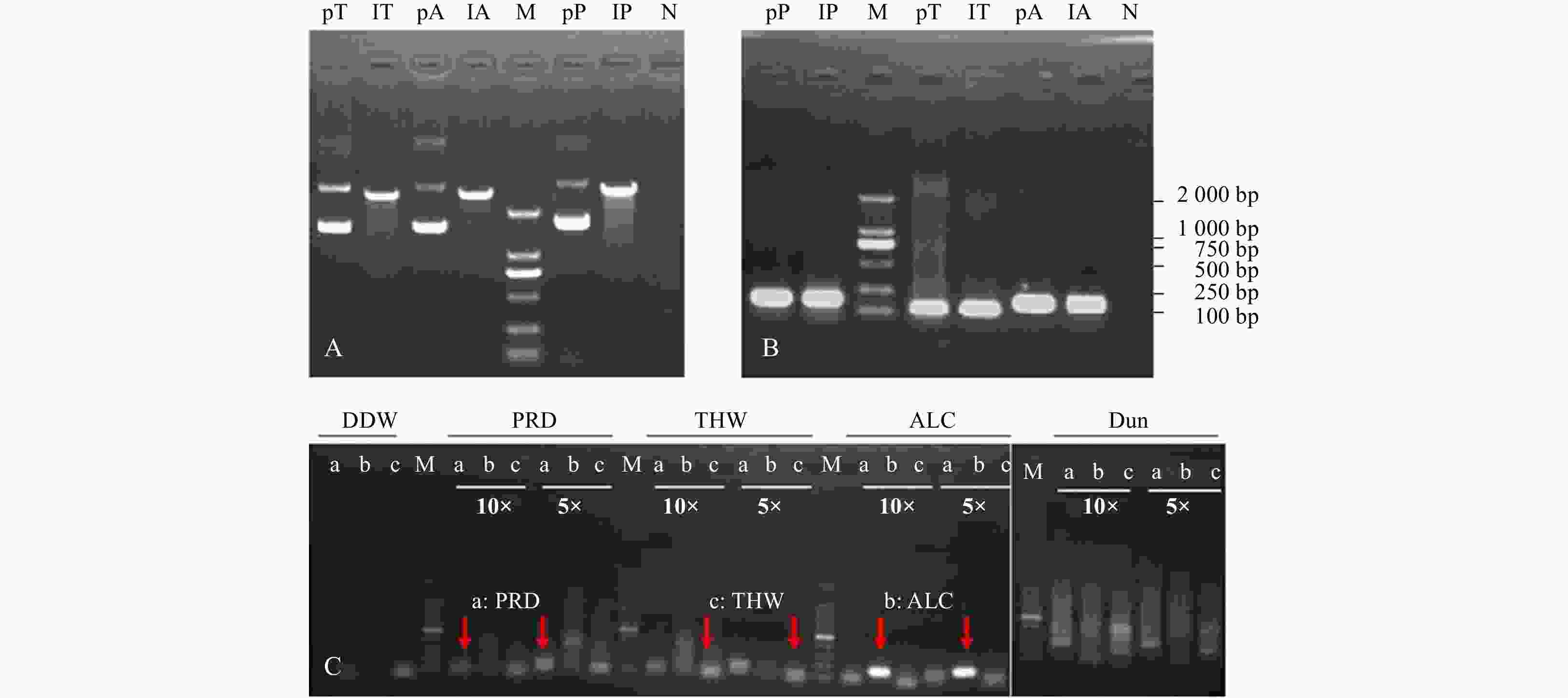
Figure 1. Extraction and linearization of plasmids inserted target 18S rDNA fragment (A), validation of the specificy primers in RT-PCR assay with plasmids above (B) and extracted DNA from fixed copepod (C) as template. pT, lT, pA, lA, pP, lP in A and B represent the templates from plasmid, linear plasmid inserted with 18S rDNA fragment of Thalassiosira weissflogii, Alexandrium catenella, and Prorocentrum shikokuense, respectively. 10× and 5× indicate the dilution time of template. a, b, and c in C display the PCR product amplified with species-specific primers of PRD, ALC, and THW, respectively. M: DNA marker 2000. N: negative. DDW: double distilled water; PRD: P. shikokuense; THW: T. weissflogii; ALC: A. catenella; Dun: Dunaliella sp. (as the control group); pT, lT: plasmid and linearization of 18S rDNA of T. weissflogii; pA, lA: plasmid and linearization of 18S rDNA of A. catenella. pP, lP: plasmid and linearization of 18S rDNA of P. shikokuense.
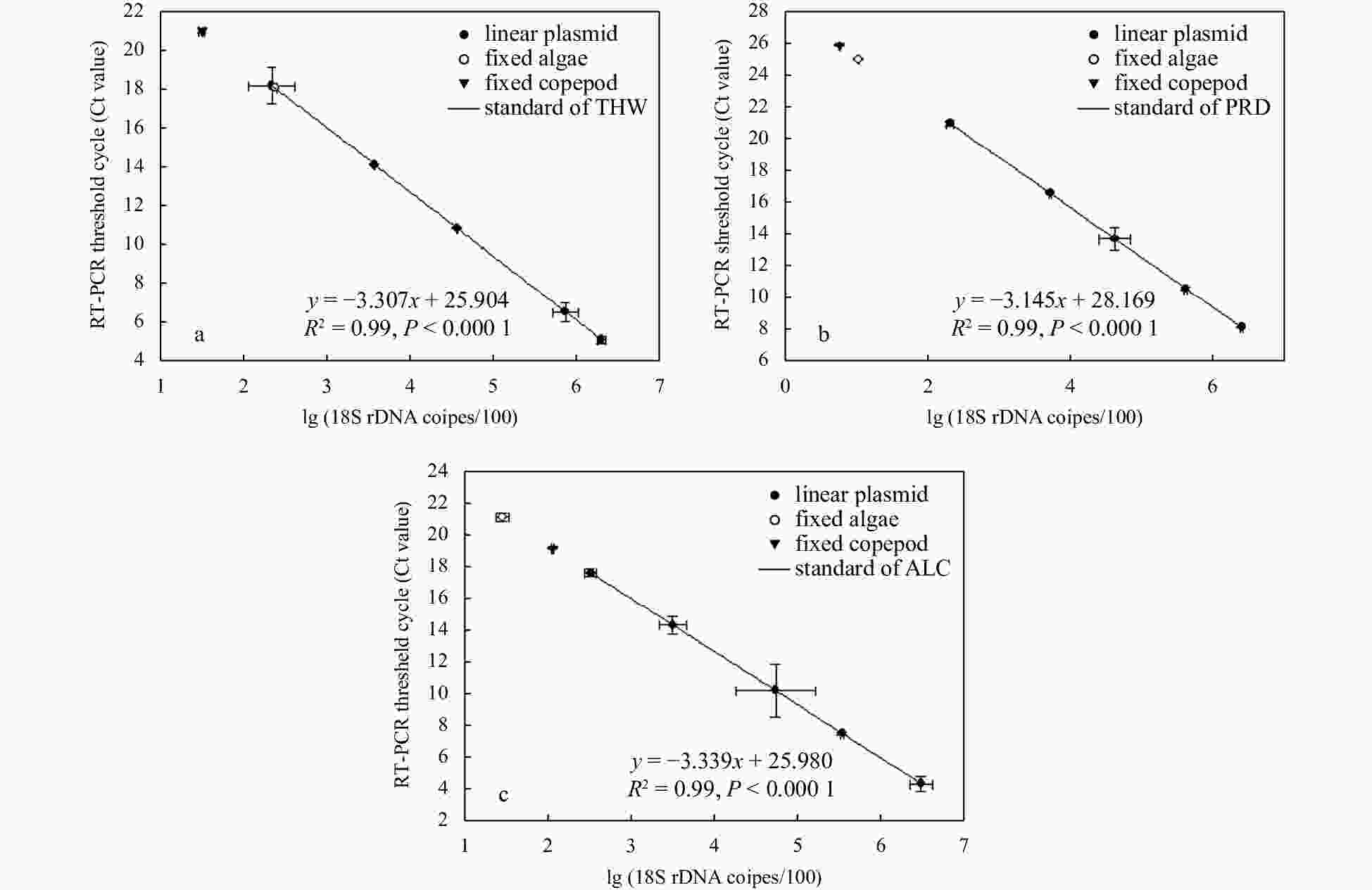
Figure 2. Quantification of microalgae 18S rDNA gene copies in fixed algae (algal cells left after feeding) and copepods based on the standards of three microalgae: Thalassiosira weissflogii (THW) (a), Prorocentrum shikokuense (PRD) (b), and Alexandrium catenella (ALC) (c). Linear plasmids inserted with target 18S rDNA fragment are used as templates to make quantitative standards for estimating cells remaining after the feeding experiment and in the copepod.
Table 1. Species-specific primers used in this study
Algae species Primer forward (5'–3') Primer reverse (5'–3') Amplicon size/bp GenBank Acc. No. Thalassiosira weissflogii ATTGGAGGGCAAGTCTGGTG GGTGTGAGACTGGTCGCTCCT 121 KF733529.1 Prorocentrum shikokuense ATTGGAGGGCAAGTCTGGTG CAGACACGTTCTCCAAGAAGA 170 AY551272.1 Alexandrium catenella ATTGGAGGGCAAGTCTGGTG CAGTTGTGCTTTCAAGATAAAT 170 AJ535392.1 Table 2. Ingestion rate of Acartia erythraea based on cell counting
Algae species Cell density before/
(cells·mL−1)Cell density after/(cells·mL−1) Copepod individuals Incubation time/
minIngestion rate/
(cells·ind·−1·h−1)Thalassiosira weissflogii 1.45 (± 0.09)×104 1.04 (± 0.07)×104 30 15 4.31 (± 1.61)×103 Prorocentrum shikokuense 1.05 (± 0.07)×103 0.88 (± 0.11)×102 30 10 1.53 (± 0.13)×103 Alexandrium catenella 2.08 (± 0.13)×103 1.37 (± 0.11)×103 30 8 1.43 (± 0.04)×103 Note: before and after the feeding experiment. -
Boersma M, Wesche A, Hirche H J. 2014. Predation of calanoid copepods on their own and other copepods’ offspring. Marine Biology, 161(4): 733–743. doi: 10.1007/s00227-013-2373-7 Calbet A, Carlotti F, Gaudy R. 2007. The feeding ecology of the copepod Centropages typicus (Kröyer). Progress in Oceanography, 72(2−3): 137–150. doi: 10.1016/j.pocean.2007.01.003 Chen Mianrun, Liu Hongbin, Chen Bingzhang. 2012. Effects of dietary essential fatty acids on reproduction rates of a subtropical calanoid copepod, Acartia erythraea. Marine Ecology Progress Series, 455: 95–110. doi: 10.3354/meps09685 Chen Mianrun, Liu Hongbin, Li Hoitung. 2013. Effect of mesozooplankton feeding selectivity on the dynamics of algae in the presence of intermediate grazers—a laboratory simulation. Marine Ecology Progress Series, 486: 47–58. doi: 10.3354/meps10393 Chikaraishi Y, Ogawa N O, Kashiyama Y, et al. 2009. Determination of aquatic food-web structure based on compound-specific nitrogen isotopic composition of amino acids. Limnology and Oceanography, 7(11): 740–750. doi: 10.4319/lom.2009.7.740 Cleary A C, Durbin E G, Rynearson T A, et al. 2016. Feeding by Pseudocalanus copepods in the Bering Sea: trophic linkages and a potential mechanism of niche partitioning. Deep-Sea Research Part II: Topical Studies in Oceanography, 134: 181–189. doi: 10.1016/j.dsr2.2015.04.001 Conroy B J, Steinberg D K, Song B, et al. 2017. Mesozooplankton graze on cyanobacteria in the Amazon River Plume and Western Tropical North Atlantic. Frontiers in Microbiology, 8: 1436. doi: 10.3389/fmicb.2017.01436 DeMott W R, Watson M D. 1991. Remote detection of algae by copepods: responses to algal size, odors and motility. Journal of Plankton Research, 13(6): 1203–1222. doi: 10.1093/plankt/13.6.1203 De Souza Santos L P, Castel J. 2013. Comparison of four methods to estimate meiobenthic copepod Amonardia normani ingestion rates. Marine Biology, 160(9): 2395–2404. doi: 10.1007/s00227-013-2234-4 Durbin E G, Casas M C, Rynearson T A. 2012. Copepod feeding and digestion rates using prey DNA and qPCR. Journal of Plankton Research, 34(1): 72–82. doi: 10.1093/plankt/fbr082 Ekmann K S, Dalsgaard J, Holm J, et al. 2013. Effects of dietary energy density and digestible protein: energy ratio on de novo lipid synthesis from dietary protein in gilthead sea bream (Sparus aurata) quantified with stable isotopes. British Journal of Nutrition, 110(10): 1771–1781. doi: 10.1017/S0007114513001281 Fields D M, Durif C M F, Bjelland R M, et al. 2011. Grazing rates of Calanus finmarchicus on Thalassiosira weissflogii cultured under different levels of ultraviolet radiation. PLoS ONE, 6(10): e26333. doi: 10.1371/journal.pone.0026333 Frost B W. 1972. Effects of size and concentration of food particles on the feeding behavior of the marine planktonic copepod Calanus pacificus. Limnology and Oceanography, 17(6): 805–815. doi: 10.4319/lo.1972.17.6.0805 Garzke J, Ismar S M H, Sommer U. 2015. Climate change affects low trophic level marine consumers: warming decreases copepod size and abundance. Oecologia, 177(3): 849–860. doi: 10.1007/s00442-014-3130-4 Godhe A, Asplund M E, Härnström K, et al. 2008. Quantification of diatom and dinoflagellate biomasses in coastal marine seawater samples by Real-Time PCR. Applied and Environmental Microbiology, 74(23): 7174–7182. doi: 10.1128/AEM.01298-08 Gonçalves A M M, Azeiteiro U M, Pardal M A, et al. 2012. Fatty acid profiling reveals seasonal and spatial shifts in zooplankton diet in a temperate estuary. Estuarine, Coastal and Shelf Science, 109: 70–80, Guo Zhiling, Liu Sheng, Hu Simin, et al. 2012. Prevalent ciliate symbiosis on copepods: high genetic diversity and wide distribution detected using small subunit ribosomal RNA gene. PLoS ONE, 7(9): e44847. doi: 10.1371/journal.pone.0044847 Hansen F C, Reckermann M, Klein Breteler W C M, et al. 1993. Phaeocystis blooming enhanced by copepod predation on protozoa: evidence from incubation experiments. Marine Ecology Progress Series, 102: 51–57. doi: 10.3354/meps102051 Hebert C E, Weseloh D V C, Gauthier L T, et al. 2009. Biochemical tracers reveal intra-specific differences in the food webs utilized by individual seabirds. Oecologia, 160(1): 15–23. doi: 10.1007/s00442-009-1285-1 Ho T W, Hwang J S, Cheung M K, et al. 2017. DNA-based study of the diet of the marine calanoid copepod Calanus sinicus. Journal of Experimental Marine Biology and Ecology, 494: 1–9. doi: 10.1016/j.jembe.2017.04.004 Hu Simin, Guo Zhiling, Li Tao, et al. 2015. Molecular analysis of in situ diets of coral reef copepods: evidence of terrestrial plant detritus as a food source in Sanya Bay, China. Journal of Plankton Research, 37(2): 363–371. doi: 10.1093/plankt/fbv014 Hu Simin, Liu Sheng, Wang Lingli, et al. 2018. Feeding response of the tropical copepod Acartia erythraea to short-term thermal stress: more animal-derived food was consumed. PeerJ, 6: e6129. doi: 10.7717/peerj.6129 Huskin I, Anadón R, Medina G, et al. 2001. Mesozooplankton distribution and copepod grazing in the subtropical Atlantic near the Azores: influence of mesoscale structures. Journal of Plankton Research, 23(7): 671–691. doi: 10.1093/plankt/23.7.671 Isari S, Antó M, Saiz E. 2013. Copepod foraging on the basis of food nutritional quality: can copepods really choose?. PLoS ONE, 8(12): e84742, Ismar S M H, Kottmann J S, Sommer U. 2018. First genetic quantification of sex- and stage-specific feeding in the ubiquitous copepod Acartia tonsa. Marine Biology, 165(2): 25. doi: 10.1007/s00227-017-3281-z Kleppel G S. 1993. On the diets of calanoid copepods. Marine Ecology Progress Series, 99: 183–195. doi: 10.3354/meps099183 LaJeunesse T C, Lambert G, Andersen R A, et al. 2005. Symbiodinium (Pyrrhophyta) genome sizes (DNA content) are smallest among dinoflagellates. Journal of Phycology, 41(4): 880–886. doi: 10.1111/j.0022-3646.2005.04231.x Liu Sheng, Li Tao, Huang Hui, et al. 2010. Feeding efficiency of a marine copepod Acartia erythraea on eight different algal diets. Acta Ecologica Sinica, 30(1): 22–26. doi: 10.1016/j.chnaes.2009.12.004 Liu Sheng, Wang Wenxiong, Huang Liangmin. 2006. Phosphorus dietary assimilation and efflux in the marine copepod Acartia erythraea. Marine Ecology Progress Series, 321: 193–202. doi: 10.3354/meps321193 Mackas D, Bohrer R. 1976. Fluorescence analysis of zooplankton gut contents and an investigation of diel feeding patterns. Journal of Experimental Marine Biology and Ecology, 25(1): 77–85. doi: 10.1016/0022-0981(76)90077-0 Nejstgaard J C, Frischer M E, Raule C L, et al. 2003. Molecular detection of algal prey in copepod guts and fecal pellets. Limnology and Oceanography: Methods, 1(1): 29–38. doi: 10.4319/lom.2003.1.29 Nejstgaard J C, Frischer M E, Simonelli P, et al. 2008. Quantitative PCR to estimate copepod feeding. Marine Biology, 153(4): 565–577. doi: 10.1007/s00227-007-0830-x Passmore A J, Jarman S N, Swadling K M, et al. 2006. DNA as a dietary biomarker in Antarctic Krill, Euphausia superba. Marine Biotechnology, 8(6): 686–696. doi: 10.1007/s10126-005-6088-8 Prokopowich C D, Gregory T R, Crease T J. 2003. The correlation between rDNA copy number and genome size in eukaryotes. Genome, 46: 48–50. doi: 10.1139/G02-103 Ray J L, Skaar K S, Simonelli P, et al. 2016. Molecular gut content analysis demonstrates that Calanus grazing on Phaeocystis pouchetii and Skeletonema marinoi is sensitive to bloom phase but not prey density. Marine Ecology Progress Series, 542: 63–77. doi: 10.3354/meps11560 Santoferrara L F. 2019. Current practice in plankton metabarcoding: optimization and error management. Journal of Plankton Research, 41(5): 571–582. doi: 10.1093/plankt/fbz041 Shin K, Jang M C, Jang P K, et al. 2003. Influence of food quality on egg production and viability of the marine planktonic copepod Acartia omorii. Progress in Oceanography, 57(3–4): 265–277. doi: 10.1016/s0079-6611(03)00101-0 Slaughter A M, Bollens S M, Bollens G R. 2006. Grazing impact of mesozooplankton in an upwelling region off northern California, 2000–2003. Deep-Sea Research Part II: Topical Studies in Oceanography, 53(25–26): 3099–3115. doi: 10.1016/j.dsr2.2006.07.005 Smith K F, Biessy L, Argyle P A, et al. 2017. Molecular identification of Gambierdiscus and Fukuyoa (Dinophyceae) from environmental samples. Marine Drugs, 15(8): 243. doi: 10.3390/md15080243 Sun Jun, Liu Dongyan. 2003. Geometric models for calculating cell biovolume and surface area for phytoplankton. Journal of Plankton Research, 25(11): 1331–1346. doi: 10.1093/plankt/fbg096 Symondson W O C. 2002. Molecular identification of prey in predator diets. Molecular Ecology, 11(4): 627–641. doi: 10.1046/j.1365-294X.2002.01471.x Troedsson C, Simonelli P, Nägele V, et al. 2009. Quantification of copepod gut content by differential length amplification quantitative PCR (dla-qPCR). Marine Biology, 156(3): 253–259. doi: 10.1007/s00227-008-1079-8 Vargas C A, Martínez R A, Cuevas L A, et al. 2007. The relative importance of microbial and classical food webs in a highly productive coastal upwelling area. Limnology and Oceanography, 52(4): 1495–1510. doi: 10.4319/lo.2007.52.4.1495 Verity P G, Smetacek V. 1996. Organism life cycles, predation, and the structure of marine pelagic ecosystems. Marine Ecology Progress Series, 130(1−3): 277–293. doi: 10.3354/meps130277 Vestheim H, Edvardsen B, Kaartvedt S. 2005. Assessing feeding of a carnivorous copepod using species-specific PCR. Marine Biology, 147(2): 381–385. doi: 10.1007/s00227-005-1590-0 Zhang Huan, Lin Senjie. 2005. Development of a cob-18S rRNA Gene Real-Time PCR Assay for Quantifying Pfiesteria shumwayae in the Natural Environment. Applied and Environmental Microbiology, 71(11): 7053–7063. doi: 10.1128/aem.71.11.7053-7063.2005 Zhu F, Massana R, Not F, et al. 2005. Mapping of picoeucaryotes in marine ecosystems with quantitative PCR of the 18S rRNA gene. FEMS Microbiology Ecology, 52(1): 79–92. doi: 10.1016/j.femsec.2004.10.006 -



 下载:
下载: