Vertical microbial profiling of water column reveals prokaryotic communities and distribution features of Antarctic Peninsula
-
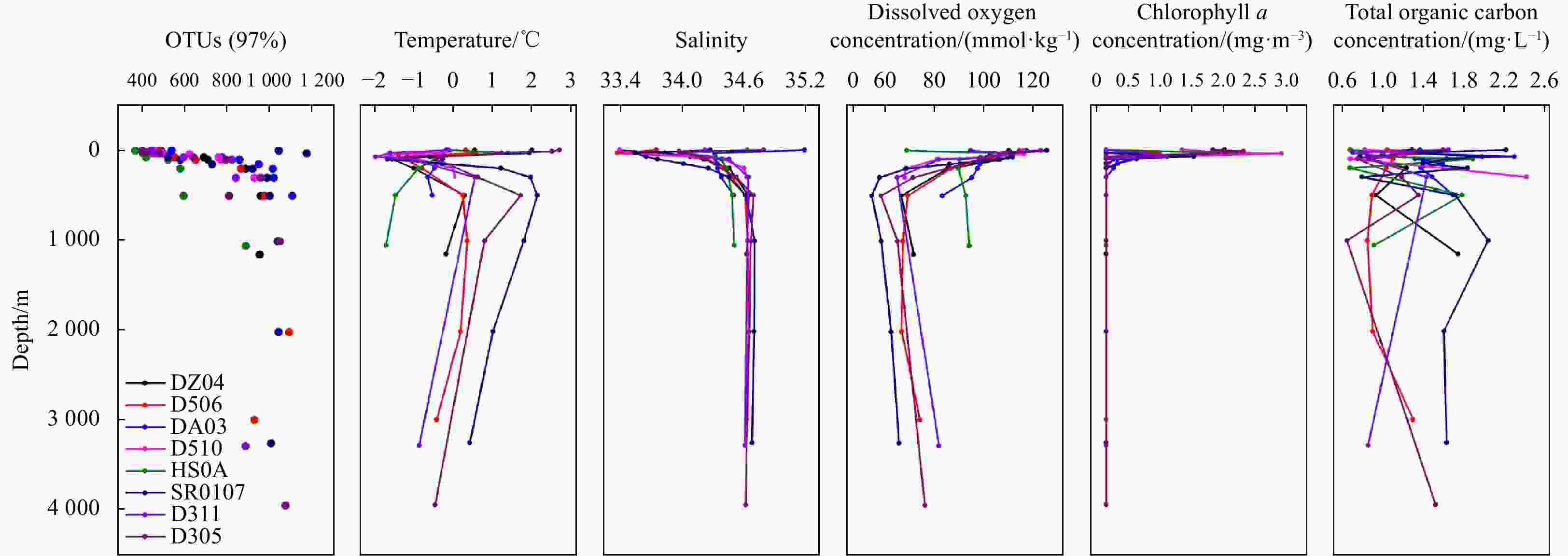
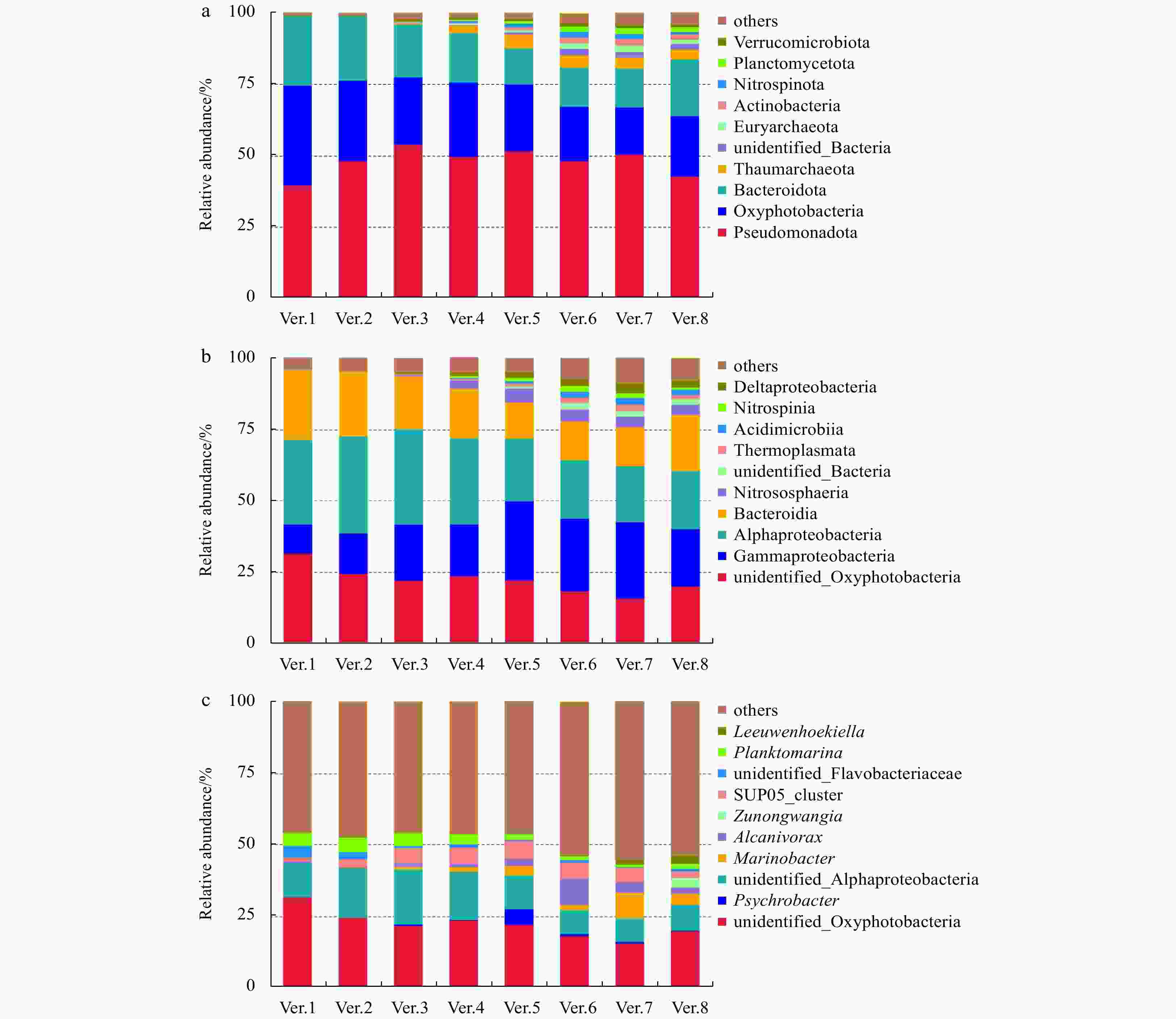
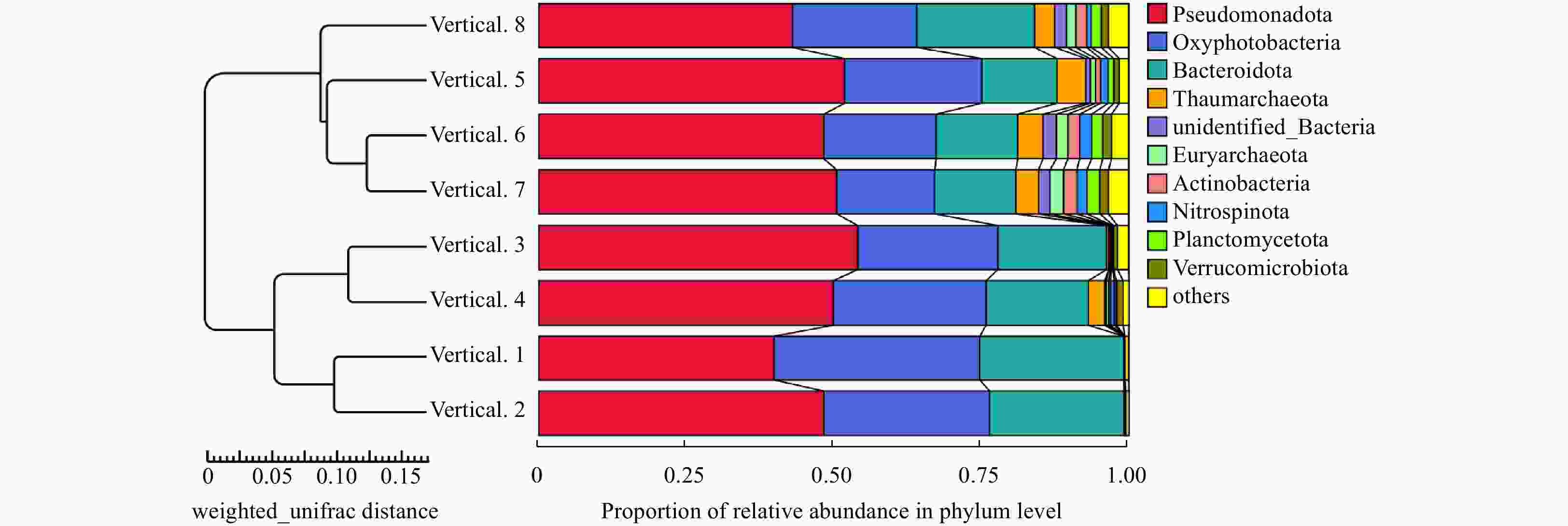
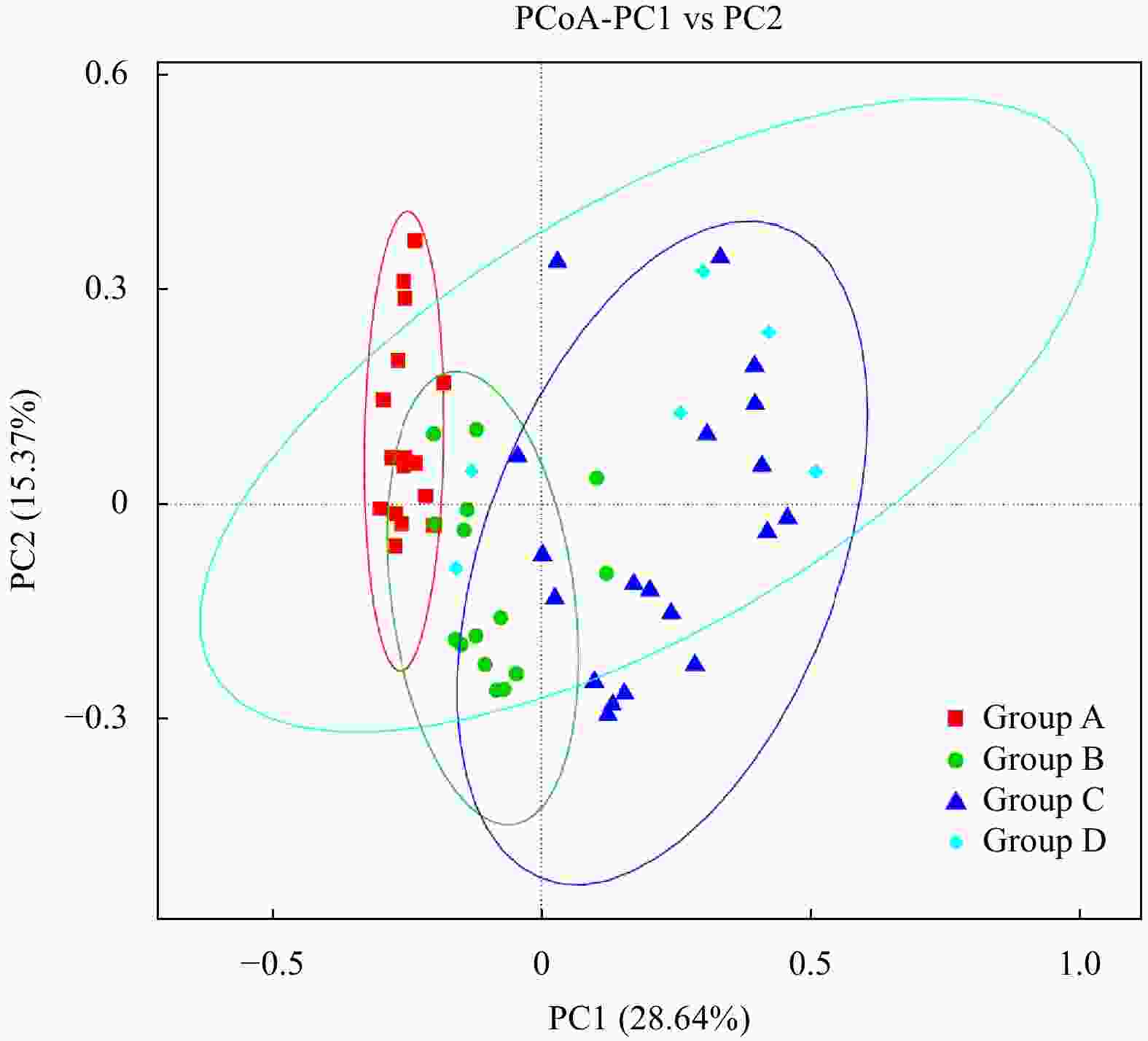
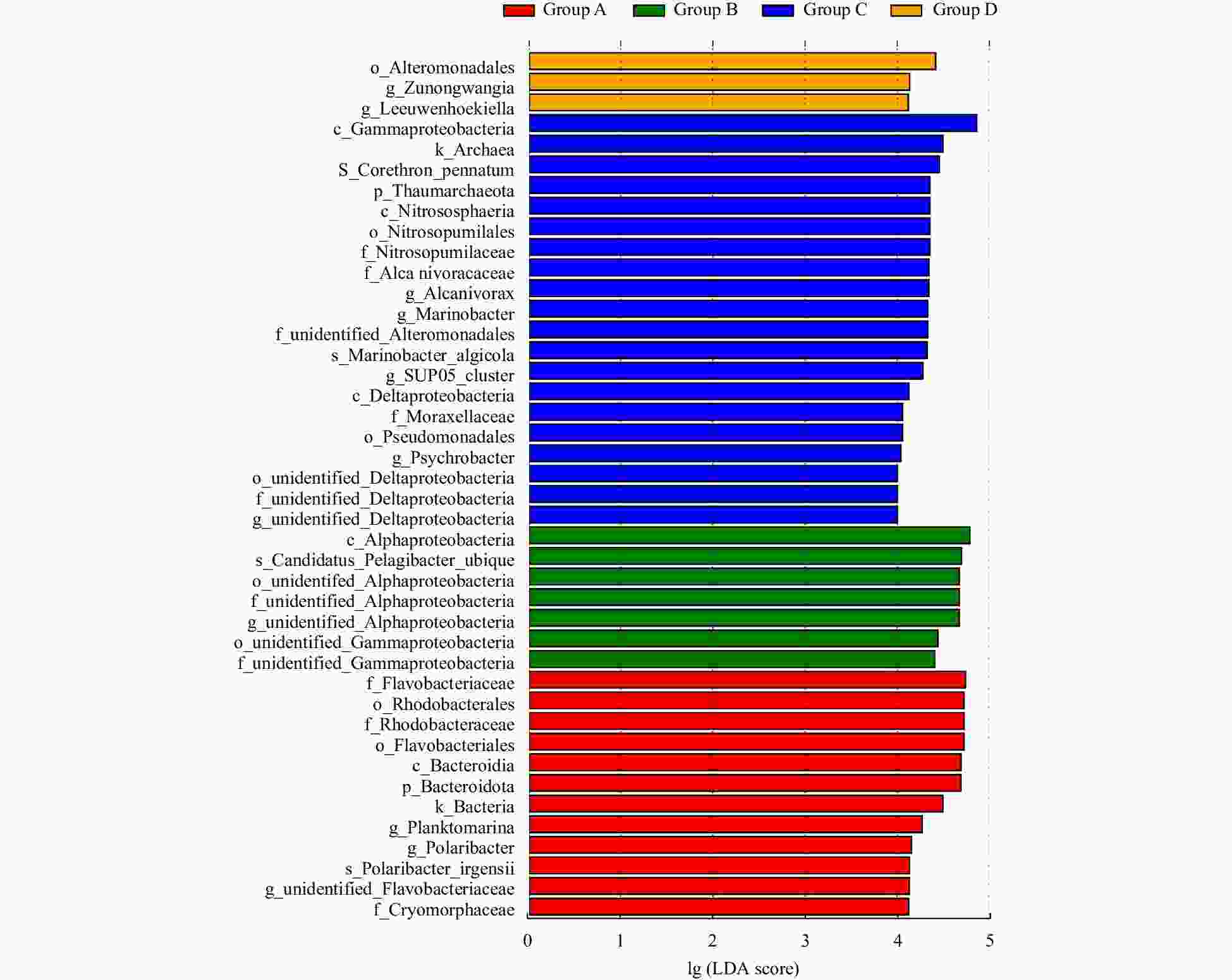
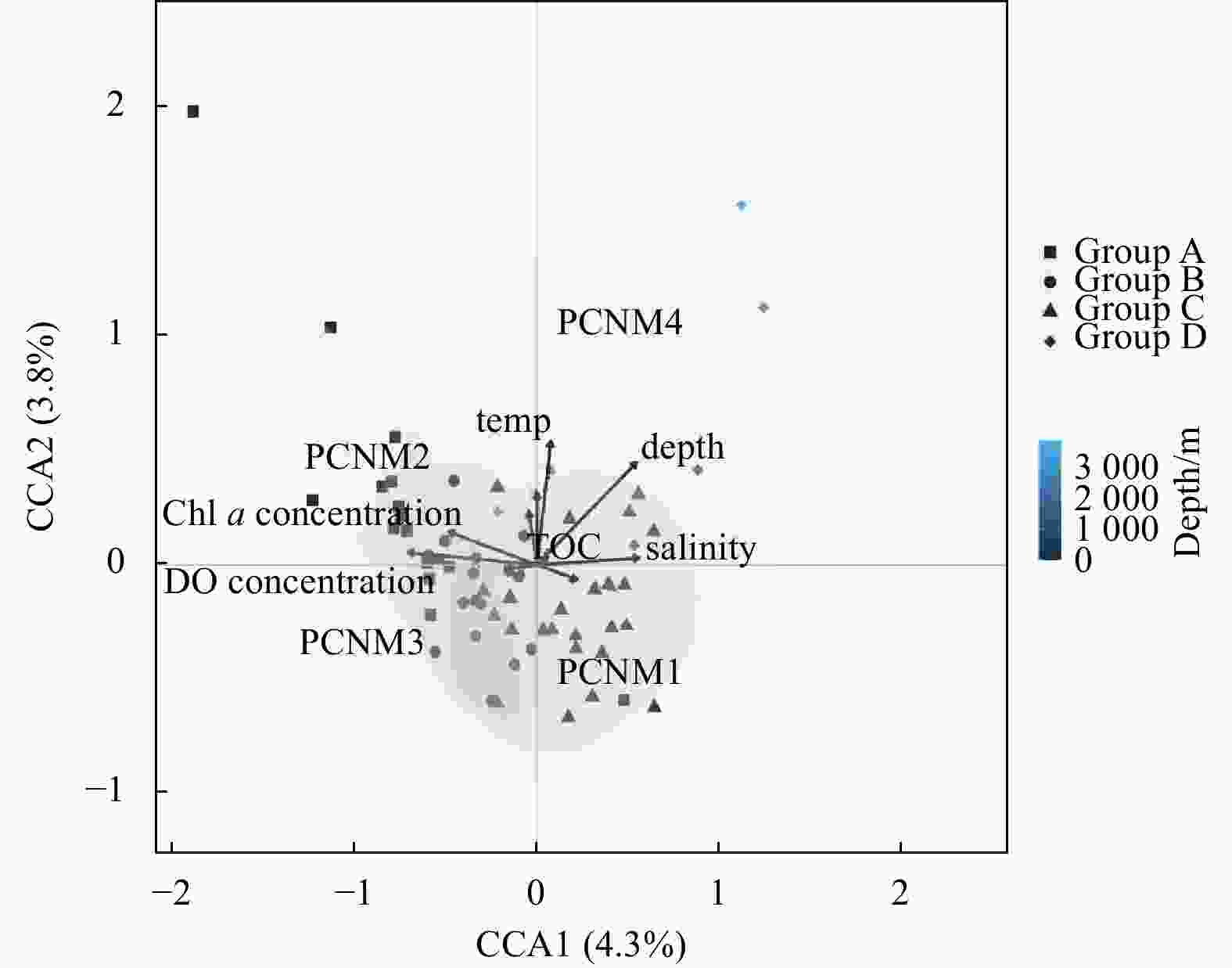
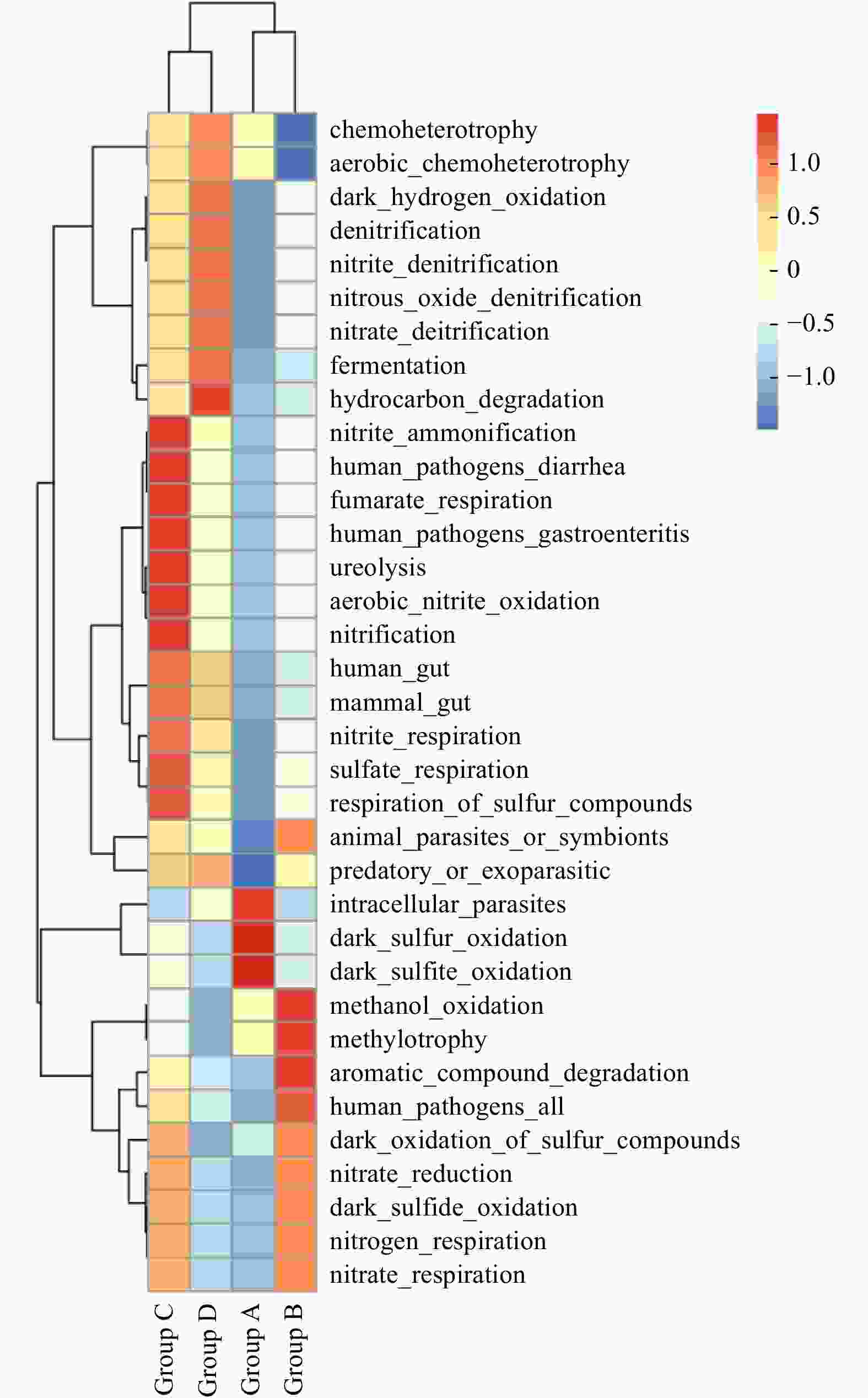
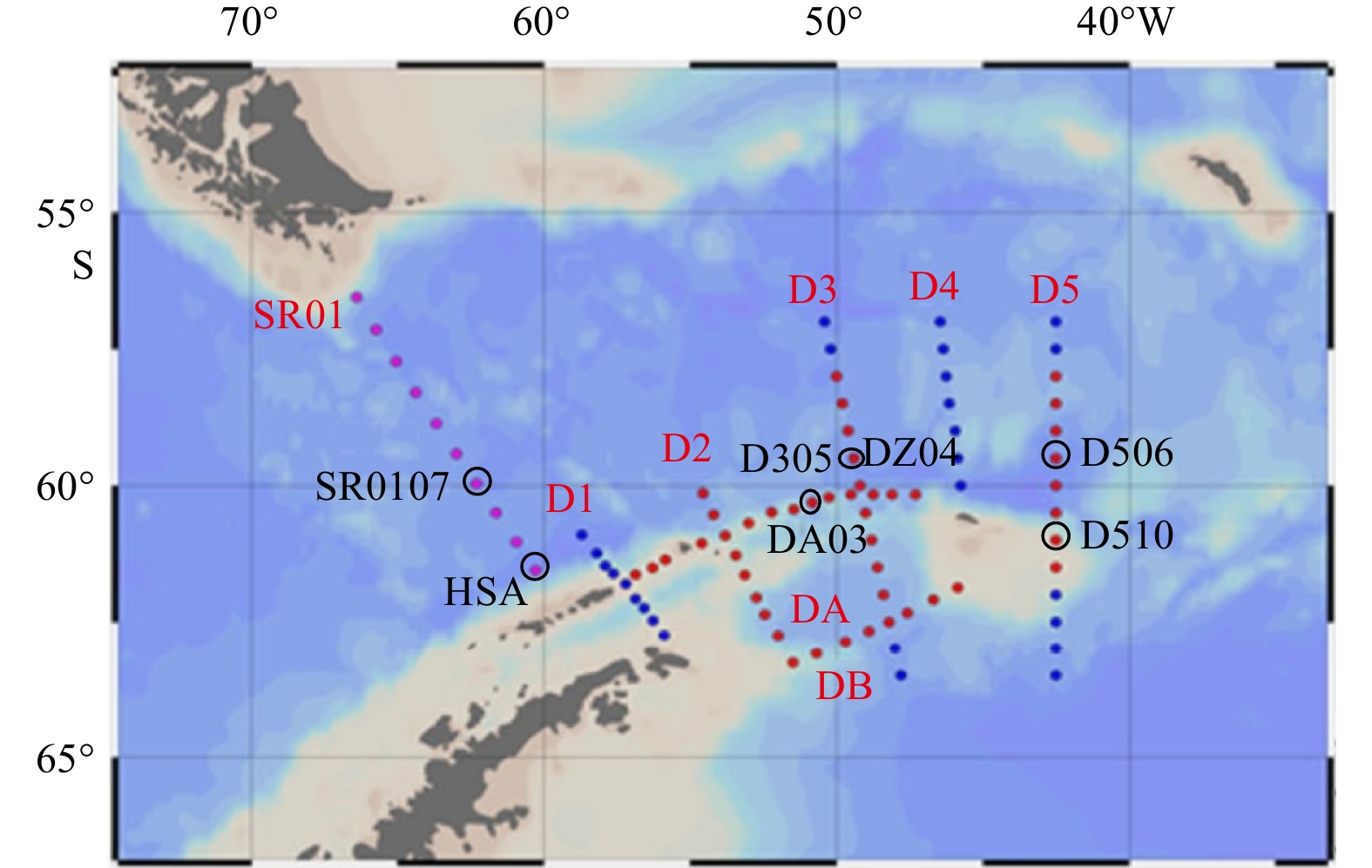
Abstract: Prokaryotic diversity and community composition in the water column of eight stations (63 samples) around the Antarctic Peninsula of the Southern Ocean were investigated. Through pyrosequencing of the V3–V4 hypervariable regions of the 16S ribosomal RNA gene, we characterized 4 720 089 valid reads representing 48 188 operational taxonomic units (OTUs, 97% similarity). The community was dominated by the phyla Pseudomonadota (original name: Proteobacteria, 47%), Oxyphotobacteria (26%), and Bacteroidota (original name: Bacteroidetes, 18%), which comprised an average of 91% of the total OTUs in all samples. The prokaryotic community composition varied vertically within the water column. Water column prokaryotic communities exhibited a clear depth profile, with higher microbial richness and higher diversity observed with increasing water depth. Cluster analysis of the community composition of water column samples exhibited a similar trend with depth. Correlation with environmental factors suggested distinct variation in prokaryotic community composition with changes in depth, salinity, temperature and dissolved oxygen levels. Functional prediction showed presence of active nitrogen, sulphur and methane metabolic cycles along the vertical transect of the studied region. These results will improve our knowledge of prokaryotic diversity and community composition at different depth of water column for better understanding of the microbial ecology and nutrient cycles in Antarctic Peninsula region of the Southern Ocean.
-
Key words:
- prokaryotic diversity /
- 16S rRNA /
- high-throughput sequencing (HTS) /
- Southern Ocean
-
Table 1. Sample sites description
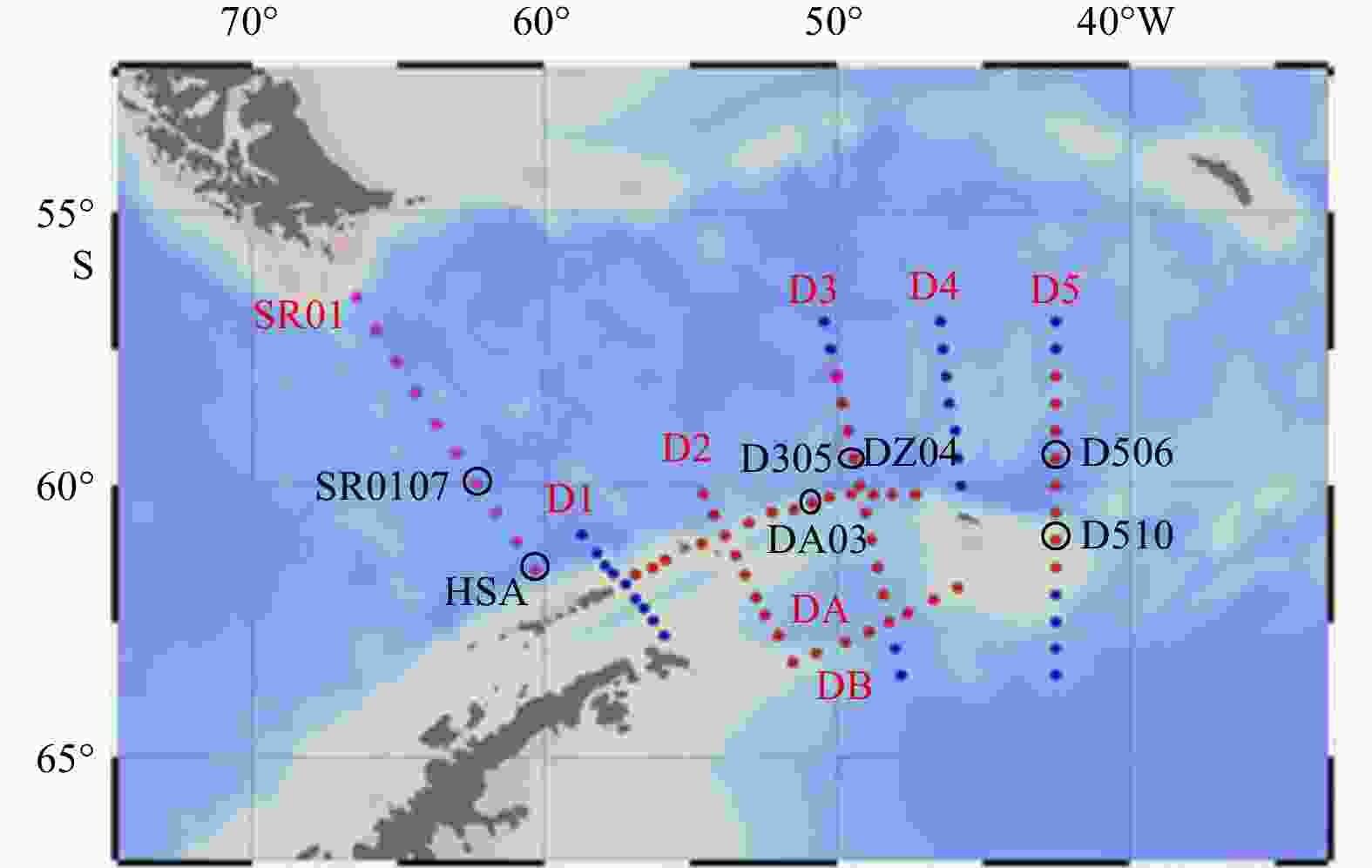
No. Station Date (UTC) Time (UTC) Latitude Longitude Depth/m 1 SR0107 2018-01-02 17:42 59°57.588' S 62°18.739'W 3 237 2 DA03 2018-01-04 9:58 60°42.002'S 53°0.073'W 456 3 D305 2018-01-09 10:11 59°0.040'S 49°36.002'W 3 922 4 D311 2018-01-10 16:35 62°0.114'S 48°23.971'W 3 266 5 D506 2018-01-20 8:01 59°29.968'S 42°30.116'W 3 001 6 D510 2018-01-21 19:34 61°17.989'S 42°29.975'W 520 7 DZ04 2018-01-29 1:35 60°40.884'S 48°56.869'W 1 147 8 HS0A 2018-02-01 17:22 62°25.914'S 58°24.620'W 1 052 Table 2. Total sequencing information
Station ID Raw_reads Clean_reads Average length/nt Unique_tag operation taxonomic unit DZ04 85 925.29 80 431.86 371.28 17 794.43 746.71 D506 81 482.22 76 914.56 370.88 22 243.22 785.66 DA03 87 184.57 76 857.00 370.71 20 270.71 888.42 D510 79 533.43 74 742.14 371.57 23 185.29 714.14 HSA 74 749.29 59 261.86 371.14 18 102.29 550.28 SR0107 80 856.36 77 193.55 370.72 22 337.00 913.54 D311 78 915.67 74 208.17 371.50 21 128.17 682.66 D305 80 973.11 77 158.89 370.66 25 446.78 741.66 Table 3. Biodiversity and richness indices of different depth water column samples
Number Depth/m Observed species Shannon Simpson Chao1 ACE Good’s coverage Vertical.1 0 408.37 5.36 0.94 537.09 537.74 0.99 Vertical.2 30 416.50 5.53 0.94 528.94 540.14 0.99 Vertical.3 75 534.75 5.91 0.95 648.74 662.71 0.99 Vertical.4 100 632.25 6.13 0.95 793.81 787.07 0.99 Vertical.5 200 769.83 6.22 0.94 962.32 982.97 0.99 Vertical.6 500 849.42 6.57 0.95 1 040.85 1 051.24 0.99 Vertical.7 1 000 874.00 6.62 0.96 1 074.29 1 078.05 0.99 Vertical.8 ≥2 000 875.00 6.49 0.96 1 092.54 1 095.79 0.99 -
Abell G C J, Bowman J P. 2005. Ecological and biogeographic relationships of class Flavobacteria in the Southern Ocean. FEMS Microbiology Ecology, 51(2): 265–277. doi: 10.1016/j.femsec.2004.09.001 Agogué H, Lamy D, Neal P R, et al. 2011. Water mass-specificity of bacterial communities in the North Atlantic revealed by massively parallel sequencing. Molecular Ecology, 20(2): 258–274. doi: 10.1111/j.1365-294X.2010.04932.x Aislabie J M, Jordan S, Barker G M. 2008. Relation between soil classification and bacterial diversity in soils of the Ross Sea region, Antarctica. Geoderma, 144(1−2): 9–20. doi: 10.1016/j.geoderma.2007.10.006 Aldunate M, De la Iglesia R, Bertagnolli A D, et al. 2018. Oxygen modulates bacterial community composition in the coastal upwelling waters off central Chile. Deep-Sea Research Part II: Topical Studies in Oceanography, 156: 68–79. doi: 10.1016/j.dsr2.2018.02.001 Beman J M, Carolan M T. 2013. Deoxygenation alters bacterial diversity and community composition in the ocean’s largest oxygen minimum zone. Nature Communications, 4: 2705. doi: 10.1038/ncomms3705 Brown M V, Lauro F M, DeMaere M Z, et al. 2012. Global biogeography of SAR11 marine bacteria. Molecular Systems Biology, 8: 595. doi: 10.1038/msb.2012.28 Brown M V, Philip G K, Bunge J A, et al. 2009. Microbial community structure in the North Pacific Ocean. The ISME Journal, 3(12): 1374–1386. doi: 10.1038/ismej.2009.86 Bryant J A, Stewart F J, Eppley J M, et al. 2012. Microbial community phylogenetic and trait diversity declines with depth in a marine oxygen minimum zone. Ecology, 93(7): 1659–1673. doi: 10.1890/11-1204.1 Caporaso J G, Kuczynski J, Stombaugh J, et al. 2010. QIIME allows analysis of high-throughput community sequencing data. Nature Methods, 7(5): 335–336. doi: 10.1038/nmeth.f.303 Chong C W, Dunn M J, Convey P, et al. 2009a. Environmental influences on bacterial diversity of soils on Signy Island, maritime Antarctic. Polar Biology, 32(11): 1571–1582. doi: 10.1007/s00300-009-0656-8 Chong C W, Goh Y S, Convey P, et al. 2013. Spatial pattern in Antarctica: what can we learn from Antarctic bacterial isolates?. Extremophiles, 17(5): 733–745 Chong C W, Tan G Y A, Wong R C S, et al. 2009b. DGGE fingerprinting of bacteria in soils from eight ecologically different sites around Casey Station, Antarctica. Polar Biology, 32(6): 853–860. doi: 10.1007/s00300-009-0585-6 Cleary D F R, de Voogd N J, Polónia A R M, et al. 2015. Composition and predictive functional analysis of bacterial communities in seawater, sediment and sponges in the Spermonde Archipelago, Indonesia. Microbial Ecology, 70(4): 889–903. doi: 10.1007/s00248-015-0632-5 DeLong E F, Preston C M, Mincer T, et al. 2006. Community genomics among stratified microbial assemblages in the ocean’s interior. Science, 311(5760): 496–503. doi: 10.1126/science.1120250 Dinasquet J, Richert I, Logares R, et al. 2017. Mixing of water masses caused by a drifting iceberg affects bacterial activity, community composition and substrate utilization capability in the Southern Ocean. Environmental Microbiology, 19(6): 2453–2467. doi: 10.1111/1462-2920.13769 Ducklow H W, Myers K M S, Erickson M, et al. 2011. Response of a summertime Antarctic marine bacterial community to glucose and ammonium enrichment. Aquatic Microbial Ecology, 64(3): 205–220. doi: 10.3354/ame01519 Esper O, Zonneveld K A F. 2002. Distribution of organic-walled dinoflagellate cysts in surface sediments of the Southern Ocean (eastern Atlantic sector) between the Subtropical Front and the Weddell Gyre. Marine Micropaleontology, 46(1−2): 177–208. doi: 10.1016/S0377-8398(02)00041-5 Fandino L B, Riemann L, Steward G F, et al. 2001. Variations in bacterial community structure during a dinoflagellate bloom analyzed by DGGE and 16S rDNA sequencing. Aquatic Microbial Ecology, 23(2): 119–130 Fierer N, Breitbart M, Nulton J, et al. 2007. Metagenomic and small-subunit rRNA analyses reveal the genetic diversity of bacteria, archaea, fungi, and viruses in soil. Applied and Environmental Microbiology, 73(21): 7059–7066. doi: 10.1128/AEM.00358-07 Francis C A, Beman J M, Kuypers M M M. 2007. New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation. The ISME Journal, 1(1): 19–27. doi: 10.1038/ismej.2007.8 Fuhrman J A, Cram J A, Needham D M. 2015. Marine microbial community dynamics and their ecological interpretation. Nature Reviews Microbiology, 13(3): 133–146. doi: 10.1038/nrmicro3417 Ghiglione J F, Galand P E, Pommier T, et al. 2012. Pole-to-pole biogeography of surface and deep marine bacterial communities. Proceedings of the National Academy of Sciences, 109(43): 17633–17638. doi: 10.1073/pnas.1208160109 Ghiglione J F, Murray A E. 2012. Pronounced summer to winter differences and higher wintertime richness in coastal Antarctic marine bacterioplankton. Environmental Microbiology, 14(3): 617–629. doi: 10.1111/j.1462-2920.2011.02601.x Giovannoni S J, Stingl U. 2005. Molecular diversity and ecology of microbial plankton. Nature, 437(7057): 343–348. doi: 10.1038/nature04158 Giovannoni S J, Vergin K L. 2012. Seasonality in ocean microbial communities. Science, 335(6069): 671–676. doi: 10.1126/science.1198078 Goffredi S K, Waren A, Orphan V J, et al. 2004. Novel forms of structural integration between microbes and a hydrothermal vent gastropod from the Indian Ocean. Applied and Environmental Microbiology, 70(5): 3082–3090. doi: 10.1128/AEM.70.5.3082-3090.2004 Grzymski J J, Riesenfeld C S, Williams T J, et al. 2012. A metagenomic assessment of winter and summer bacterioplankton from Antarctica Peninsula coastal surface waters. The ISME Journal, 6(10): 1901–1915. doi: 10.1038/ismej.2012.31 Hammer Ø, Harper D A T, Ryan P D. 2001. PAST: Palaeontological Statistics Software Package for Education and Data Analysis. Palaeontologia Electronica, 4(1): 1–9 Hartmann M, Gomez-Pereira P, Grob C, et al. 2014. Efficient CO2 fixation by surface Prochlorococcus in the Atlantic Ocean. The ISME Journal, 8(11): 2280–2289 Jamieson R E, Rogers A D, Billett D S M, et al. 2012. Patterns of marine bacterioplankton biodiversity in the surface waters of the Scotia Arc, Southern Ocean. FEMS Microbiology Ecology, 80(2): 452–468. doi: 10.1111/j.1574-6941.2012.01313.x Jayakumar A, Chang B X, Widner B, et al. 2017. Biological nitrogen fixation in the oxygen-minimum region of the eastern tropical North Pacific Ocean. The ISME Journal, 11(10): 2356–2367. doi: 10.1038/ismej.2017.97 Karlson A M L, Duberg J, Motwani N H, et al. 2015. Nitrogen fixation by cyanobacteria stimulates production in Baltic food webs. AMBIO, 44(S3): 413–426. doi: 10.1007/s13280-015-0660-x Kembel S W, Eisen J A, Pollard K S, et al. 2011. The phylogenetic diversity of metagenomes. PLoS ONE, 6(8): e23214. doi: 10.1371/journal.pone.0023214 Landa M, Blain S, Christaki U, et al. 2016. Shifts in bacterial community composition associated with increased carbon cycling in a mosaic of phytoplankton blooms. The ISME Journal, 10(1): 39–50. doi: 10.1038/ismej.2015.105 Laws E A, Falkowski P G, Smith Jr W O, et al. 2000. Temperature effects on export production in the open ocean. Global Biogeochemical Cycles, 14(4): 1231–1246. doi: 10.1029/1999GB001229 Liu Ning, Zhou Jialiang, Han Lujia, et al. 2017. Role and multi-scale characterization of bamboo biochar during poultry manure aerobic composting. Bioresource Technology, 241: 190–199. doi: 10.1016/j.biortech.2017.03.144 Loescher C R, Großkopf T, Desai F D, et al. 2014. Facets of diazotrophy in the oxygen minimum zone waters off Peru. The ISME Journal, 8(11): 2180–2192. doi: 10.1038/ismej.2014.71 Manganelli M, Malfatti F, Samo T J, et al. 2009. Major role of microbes in carbon fluxes during austral winter in the southern drake passage. PLoS ONE, 4(9): e6941. doi: 10.1371/journal.pone.0006941 Mincer T J, Church M J, Taylor L T, et al. 2007. Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific subtropical gyre. Environmental Microbiology, 9(5): 1162–1175. doi: 10.1111/j.1462-2920.2007.01239.x Murray A E, Grzymski J J. 2007. Diversity and genomics of Antarctic marine micro-organisms. Philosophical Transactions of the Royal Society B: Biological Sciences, 362(1488): 2259–2271. doi: 10.1098/rstb.2006.1944 Murray A E, Peng V, Tyler C, et al. 2011. Marine bacterioplankton biomass, activity and community structure in the vicinity of Antarctic icebergs. Deep-Sea Research Part II: Topical Studies in Oceanography, 58(11−12): 1407–1421. doi: 10.1016/j.dsr2.2010.11.021 Nakagawa S, Takai K, Inagaki F, et al. 2005. Distribution, phylogenetic diversity and physiological characteristics of epsilon-Proteobacteria in a deep-sea hydrothermal field. Environmental Microbiology, 7(10): 1619–1632. doi: 10.1111/j.1462-2920.2005.00856.x Oren A, Garrity G M. 2021. Valid publication of the names of forty-two phyla of prokaryotes. International Journal of Systematic and Evolutionary Microbiology, 71(10): 005056 Park P K. 1969. A practical handbook of seawater analysis. Fisheries Research Board of Canada Bulletin 167. J. D. H. Strickland, T. R. Parsons. The Quarterly Review of Biology , 44 (3): 327. Pommier T, Neal P R, Gasol J M, et al. 2010. Spatial patterns of bacterial richness and evenness in the NW Mediterranean Sea explored by pyrosequencing of the 16S rRNA. Aquatic Microbial Ecology, 61(3): 221–233. doi: 10.3354/ame01484 R D Core Team. 2008. R: A Language and Environment for Statistical Computing. Vienna, Austria: R Foundation for Statistical Computing. http://www.R-project.org, ISBN 3-900051-07-0"[2022-03-21/2022-03-23] Segata N, Izard J, Waldron L, et al. 2011. Metagenomic biomarker discovery and explanation. Genome Biology, 12(6): R60. doi: 10.1186/gb-2011-12-6-r60 Selje N, Simon M, Brinkhoff T. 2004. A newly discovered Roseobacter cluster in temperate and polar oceans. Nature, 427(6973): 445–448. doi: 10.1038/nature02272 Shivaji S, Kumari K, Kishore K H, et al. 2011. Vertical distribution of bacteria in a lake sediment from Antarctica by culture-independent and culture-dependent approaches. Research in Microbiology, 162(2): 191–203. doi: 10.1016/j.resmic.2010.09.020 Swan B K, Martinez-Garcia M, Preston C M, et al. 2011. Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science, 333(6047): 1296–1300. doi: 10.1126/science.1203690 The Human Microbiome Project Consortium. 2012. Structure, function and diversity of the healthy human microbiome. Nature, 486(7402): 207–214. doi: 10.1038/nature11234 Thomas F, Hehemann J H, Rebuffet E, et al. 2011. Environmental and gut Bacteroidetes: the food connection. Frontiers in Microbiology, 2: 93 Torstensson A, Dinasquet J, Chierici M, et al. 2015. Physicochemical control of bacterial and protist community composition and diversity in Antarctic sea ice. Environmental Microbiology, 17(10): 3869–3881. doi: 10.1111/1462-2920.12865 Treusch A H, Vergin K L, Finlay L A, et al. 2009. Seasonality and vertical structure of microbial communities in an ocean gyre. The ISME Journal, 3(10): 1148–1163. doi: 10.1038/ismej.2009.60 Venkatachalam S, Matcher G F, Lamont T, et al. 2019. Influence of oceanographic variability on near-shore microbial communities of the sub-Antarctic Prince Edward Islands. Limnology and Oceanography, 64(1): 258–271. doi: 10.1002/lno.11035 Venter J C, Remington K, Heidelberg J F, et al. 2004. Environmental genome shotgun sequencing of the Sargasso Sea. Science, 304(5667): 66–74. doi: 10.1126/science.1093857 Walsh E A, Kirkpatrick J B, Rutherford S D, et al. 2016. Bacterial diversity and community composition from seasurface to subseafloor. The ISME Journal, 10(4): 979–989. doi: 10.1038/ismej.2015.175 Walsh E A, Smith D C, Sogin M L, et al. 2015. Bacterial and archaeal biogeography of the deep chlorophyll maximum in the South Pacific Gyre. Aquatic Microbial Ecology, 75(1): 1–13. doi: 10.3354/ame01746 Ward B B. 1996. Nitrification and ammonification in aquatic systems. Life Support and Biosphere Science: International Journal of Earth Space, 3: 25–29 Ward P, Whitehouse M, Brandon M, et al. 2003. Mesozooplankton community structure across the Antarctic Circumpolar Current to the north of South Georgia: Southern Ocean. Marine Biology, 143(1): 121–130. doi: 10.1007/s00227-003-1019-6 West N J, Obernosterer I, Zemb O, et al. 2008. Major differences of bacterial diversity and activity inside and outside of a natural iron-fertilized phytoplankton bloom in the Southern Ocean. Environmental Microbiology, 10(3): 738–756. doi: 10.1111/j.1462-2920.2007.01497.x Wilkins D, Lauro F M, Williams T J, et al. 2013a. Biogeographic partitioning of Southern Ocean microorganisms revealed by metagenomics. Environmental Microbiology, 15(5): 1318–1333. doi: 10.1111/1574-6976.12007 Wilkins D, van Sebille E, Rintoul S R, et al. 2013b. Advection shapes Southern Ocean microbial assemblages independent of distance and environment effects. Nature Communications, 4: 2457. doi: 10.1038/ncomms3457 Wilkins D, Yau S, Williams T J, et al. 2012. Key microbial drivers in Antarctic aquatic environments. FEMS Microbiology Reviews, 37(3): 303–335 Williams T J, Long E, Evans F, et al. 2012. A metaproteomic assessment of winter and summer bacterioplankton from Antarctic Peninsula coastal surface waters. The ISME Journal, 6(10): 1883–1900. doi: 10.1038/ismej.2012.28 Wright J J, Konwar K M, Hallam S J. 2012. Microbial ecology of expanding oxygen minimum zones. Nature Reviews Microbiology, 10(6): 381–394. doi: 10.1038/nrmicro2778 Yergeau E, Bokhorst S, Huiskes A H L, et al. 2007a. Size and structure of bacterial, fungal and nematode communities along an Antarctic environmental gradient. FEMS Microbiology Ecology, 59(2): 436–451 Yergeau E, Bokhorst S, Kang Sanghoon, et al. 2012. Shifts in soil microorganisms in response to warming are consistent across a range of Antarctic environments. The ISME Journal, 6(3): 692–702. doi: 10.1038/ismej.2011.124 Yergeau E, Kang Sanghoon, He Zhili, et al. 2007b. Functional microarray analysis of nitrogen and carbon cycling genes across an Antarctic latitudinal transect. The ISME Journal, 1(2): 163–179. doi: 10.1038/ismej.2007.24 Yergeau E, Newsham K K, Pearce D A, et al. 2007c. Patterns of bacterial diversity across a range of Antarctic terrestrial habitats. Environmental Microbiology, 9(11): 2670–2682. doi: 10.1111/j.1462-2920.2007.01379.x Zinger L, Amaral-Zettler L A, Fuhrman J A, et al. 2011. Global patterns of bacterial beta-diversity in seafloor and seawater ecosystems. PLoS ONE, 6(9): e24570. doi: 10.1371/journal.pone.0024570 -



 下载:
下载: